You are browsing environment: FUNGIDB
CAZyme Information: QSL66238.1
You are here: Home > Sequence: QSL66238.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Pneumocystis wakefieldiae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Pneumocystidomycetes; ; Pneumocystidaceae; Pneumocystis; Pneumocystis wakefieldiae | |||||||||||
| CAZyme ID | QSL66238.1 | |||||||||||
| CAZy Family | GT39 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 371059; End:374986 Strand: - | |||||||||||
Full Sequence Download help
| MDLNIKAERM LDLIKIFELK DSFTGIMDMN GGIHMKNREQ NGYCIQQAPS FRQNLVHNHQ | 60 |
| YCSYSKENSI ISIDELAELN LNSSVNAVAH DIGIASDSPF PVCRNVNLSY TDDFYSIDVT | 120 |
| PIAIENSRVI QSMSMPIRMD DYNSSQIYSK DSFKSFFDSN SENMIYKKIK NESSVFIEEP | 180 |
| FMSFNVFDDK KLLEKYRKRR ESHNAVERRR RDNINEKIQE LAALVLRNPF KDKIKLNMDS | 240 |
| RDKGYCDFKL NKGEILQQSV NYIQCLQNYV NELNSRSKLL EDEVIKLGGD INFTNEIKPV | 300 |
| SYIPLNIENM DSNGCDDNSF NKKLRIYLEV CNRVGGIYSV LKSKAPVTSL EYPYDRYIAI | 360 |
| GKFTQSAINF EVEQLSTEDQ PTLHKTLQYM KNRGISYTYG RWLIDGAPRV LLFDIGNAFH | 420 |
| WLNEWKGDLW NTTGIPSPPD DSETNDAIAF GYIVAWFLGE FISHEYERAV VVHFHEWLAG | 480 |
| VALPLCRKRR LDLTTIFTTH ATILGRYLCA DNADFYNNLQ YFDVDYEAGK RGIYHRYCIE | 540 |
| RAAAHSADVF TTVSHITAYE SEHLLKRKPD GVLPNGLNVV KFAAVHEFQN LHATFKQKIN | 600 |
| EFVRGHFYGH NDFDLDNTLY FFISGRYEYR NKGVDLYIES LARLNQRMKA SDSKMTVIAF | 660 |
| IIMPAVTHSY TVEALKGQAV MKSLQNTIEH IQKGIGQRIY EKCCRHSKDF DEKFIDTSSL | 720 |
| LTDQDKILLK RRVFALKRSI LPPIVTHNMA DNNDPILTHI RRVQLFNSDE DRVKIIFHPE | 780 |
| FLNSNNPILP LDYDDFVRGC HLGVFPSYYE PWGYTSAEAV VMGVPSITTN ISGFGCYMEN | 840 |
| IIDNPSDYGV YIVDRRMKSV DESINQLTDI MYDFTTKTRR QRINQRNRCE RLSDFLDWKI | 900 |
| MGLEYIKARL LALRRAYPNS FDTDLIDKNI DLALGTLIKV PHPLSTPTSP KYRNGLLSIH | 960 |
| DLDDIQNI | 968 |
Enzyme Prediction help
| EC | 2.4.1.11:48 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT3 | 328 | 965 | 1.5e-281 | 0.9905808477237049 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 340824 | GT3_GSY2-like | 0.0 | 329 | 915 | 10 | 590 | glycogen synthase GSY2 and similar proteins. Glycogen synthase, which is most closely related to the GT3 family of glycosyltransferases, catalyzes the transfer of a glucose molecule from UDP-glucose to a terminal branch of a glycogen molecule, a rate-limit step of glycogen biosynthesis. GSY2, the member of this family in S. cerevisiae, has been shown to possess glycogen synthase activity. |
| 399009 | Glycogen_syn | 0.0 | 329 | 964 | 5 | 635 | Glycogen synthase. This family consists of the eukaryotic glycogen synthase proteins GYS1, GYS2 and GYS3. Glycogen synthase (GS) is the enzyme responsible for the synthesis of -1,4-linked glucose chains in glycogen. It is the rate limiting enzyme in the synthesis of the polysaccharide, and its activity is highly regulated through phosphorylation at multiple sites and also by allosteric effectors, mainly glucose 6-phosphate (G6P). |
| 381393 | bHLHzip_USF_MITF | 2.73e-20 | 202 | 272 | 1 | 58 | basic Helix-Loop-Helix-zipper (bHLHzip) domain found in USF/MITF family. The USF (upstream stimulatory factor)/MITF (microphthalmia-associated transcription factor) family includes two bHLHzip transcription factor subfamilies. USFs are ubiquitously expressed and key regulators of a wide number of gene regulation networks, including the stress and immune responses, cell cycle and proliferation, lipid and glucid metabolism. USFs recruit chromatin remodeling enzymes and interact with co-activators and the members of the transcription pre-initiation complex. USFs interact with high affinity to E-box regulatory elements. The MITF (also known as microphthalmia-TFE, or MiT) subfamily comprises four genes in mammals (MITF, TFE3, TFEB, and TFEC); each gene has different functions. MITF is involved in neural crest melanocytes development as well as the pigmented retinal epithelium. TFEB is required for vascularization of the mouse placenta. TFE3 is involved in B cell function. TFEC regulates gene expression in macrophages. The MITF subfamily proteins can form homodimers or heterodimers with each other but not with other bHLH or bHLHzip proteins. |
| 381404 | bHLHzip_scCBP1 | 1.13e-15 | 194 | 281 | 2 | 71 | basic Helix-Loop-Helix-zipper (bHLHzip) domain found in Saccharomyces cerevisiae centromere-binding protein 1 (CBP-1) and similar proteins. CBP-1, also termed centromere promoter factor 1 (CPF1), or centromere-binding factor 1 (CBF1), is a bHLHzip protein that is required for chromosome stability and methionine prototrophy. It binds as a homodimer to the centromere DNA elements I (CDEI, GTCACATG) region of the centromere that is required for optimal centromere function. |
| 381403 | bHLHzip_MITF_like | 1.01e-13 | 195 | 273 | 1 | 64 | basic Helix-Loop-Helix-zipper (bHLHzip) domain found in the microphthalmia-associated transcription factor family (MITF) family. The MITF (also known as microphthalmia-TFE, or MiT) family is a small family that contain a basic helix loop helix domain associated with a leucine zipper (bHLHZip). The MITF family comprises four genes in mammals (MITF, TFE3, TFEB, and TFEC); each gene has different functions. MITF is involved in neural crest melanocytes development as well as the pigmented retinal epithelium. TFEB is required for vascularization of the mouse placenta. TFE3 is involved in B cell function. TFEC regulates gene expression in macrophages. The MITF family can form homodimers or heterodimers with each other but not with other bHLH or bHLHzip proteins. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QSL66238.1|GT3 | 0.0 | 1 | 968 | 1 | 968 |
| BCS25120.1|GT3 | 0.0 | 326 | 968 | 23 | 660 |
| EAA58813.1|GT3 | 0.0 | 326 | 968 | 20 | 657 |
| CAK44379.1|GT3 | 0.0 | 329 | 968 | 26 | 660 |
| BCR93830.1|GT3 | 0.0 | 329 | 968 | 26 | 660 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6U77_A | 5.36e-288 | 329 | 951 | 16 | 657 | yGsy2p in complex with small molecule [Saccharomyces cerevisiae S288C],6U77_B yGsy2p in complex with small molecule [Saccharomyces cerevisiae S288C],6U77_C yGsy2p in complex with small molecule [Saccharomyces cerevisiae S288C],6U77_D yGsy2p in complex with small molecule [Saccharomyces cerevisiae S288C] |
| 5UX7_A | 9.14e-288 | 329 | 951 | 36 | 677 | Activated state yeast Glycogen Synthase in complex with UDP-xylose [Saccharomyces cerevisiae S288C],5UX7_B Activated state yeast Glycogen Synthase in complex with UDP-xylose [Saccharomyces cerevisiae S288C],5UX7_C Activated state yeast Glycogen Synthase in complex with UDP-xylose [Saccharomyces cerevisiae S288C],5UX7_D Activated state yeast Glycogen Synthase in complex with UDP-xylose [Saccharomyces cerevisiae S288C] |
| 5SUK_A | 1.09e-287 | 329 | 951 | 36 | 677 | G6P bound activated state of yeast glycogen synthase 2 [Saccharomyces cerevisiae S288C],5SUK_B G6P bound activated state of yeast glycogen synthase 2 [Saccharomyces cerevisiae S288C],5SUK_C G6P bound activated state of yeast glycogen synthase 2 [Saccharomyces cerevisiae S288C],5SUK_D G6P bound activated state of yeast glycogen synthase 2 [Saccharomyces cerevisiae S288C] |
| 5UW0_A | 2.60e-287 | 329 | 951 | 36 | 677 | Chain A, Glycogen [starch] synthase isoform 2 [Saccharomyces cerevisiae S288C],5UW0_B Chain B, Glycogen [starch] synthase isoform 2 [Saccharomyces cerevisiae S288C],5UW0_C Chain C, Glycogen [starch] synthase isoform 2 [Saccharomyces cerevisiae S288C],5UW0_D Chain D, Glycogen [starch] synthase isoform 2 [Saccharomyces cerevisiae S288C],5UW1_A Activated state yGsy2p in complex with UDP-galactose [Saccharomyces cerevisiae S288C],5UW1_B Activated state yGsy2p in complex with UDP-galactose [Saccharomyces cerevisiae S288C],5UW1_C Activated state yGsy2p in complex with UDP-galactose [Saccharomyces cerevisiae S288C],5UW1_D Activated state yGsy2p in complex with UDP-galactose [Saccharomyces cerevisiae S288C],5UW4_A Chain A, Glycogen [starch] synthase isoform 2 [Saccharomyces cerevisiae S288C],5UW4_B Chain B, Glycogen [starch] synthase isoform 2 [Saccharomyces cerevisiae S288C],5UW4_C Chain C, Glycogen [starch] synthase isoform 2 [Saccharomyces cerevisiae S288C],5UW4_D Chain D, Glycogen [starch] synthase isoform 2 [Saccharomyces cerevisiae S288C],5VNC_A Chain A, Glycogen [starch] synthase isoform 2 [Saccharomyces cerevisiae S288C],5VNC_B Chain B, Glycogen [starch] synthase isoform 2 [Saccharomyces cerevisiae S288C],5VNC_C Chain C, Glycogen [starch] synthase isoform 2 [Saccharomyces cerevisiae S288C],5VNC_D Chain D, Glycogen [starch] synthase isoform 2 [Saccharomyces cerevisiae S288C] |
| 4KQM_A | 2.99e-287 | 329 | 951 | 35 | 676 | Crystal structure of yeast glycogen synthase E169Q mutant in complex with glucose and UDP [Saccharomyces cerevisiae FostersO],4KQM_B Crystal structure of yeast glycogen synthase E169Q mutant in complex with glucose and UDP [Saccharomyces cerevisiae FostersO],4KQM_C Crystal structure of yeast glycogen synthase E169Q mutant in complex with glucose and UDP [Saccharomyces cerevisiae FostersO],4KQM_D Crystal structure of yeast glycogen synthase E169Q mutant in complex with glucose and UDP [Saccharomyces cerevisiae FostersO] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| sp|O93869|GYS_NEUCR | 1.51e-312 | 329 | 966 | 22 | 653 | Glycogen [starch] synthase OS=Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) OX=367110 GN=gsy-1 PE=2 SV=2 |
| sp|P23337|GYS1_YEAST | 1.47e-295 | 329 | 954 | 16 | 660 | Glycogen [starch] synthase isoform 1 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=GSY1 PE=1 SV=3 |
| sp|P27472|GYS2_YEAST | 1.94e-287 | 329 | 951 | 16 | 657 | Glycogen [starch] synthase isoform 2 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=GSY2 PE=1 SV=3 |
| sp|P13807|GYS1_HUMAN | 4.93e-246 | 329 | 950 | 35 | 646 | Glycogen [starch] synthase, muscle OS=Homo sapiens OX=9606 GN=GYS1 PE=1 SV=2 |
| sp|A2RRU1|GYS1_RAT | 1.02e-245 | 329 | 950 | 35 | 646 | Glycogen [starch] synthase, muscle OS=Rattus norvegicus OX=10116 GN=Gys1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
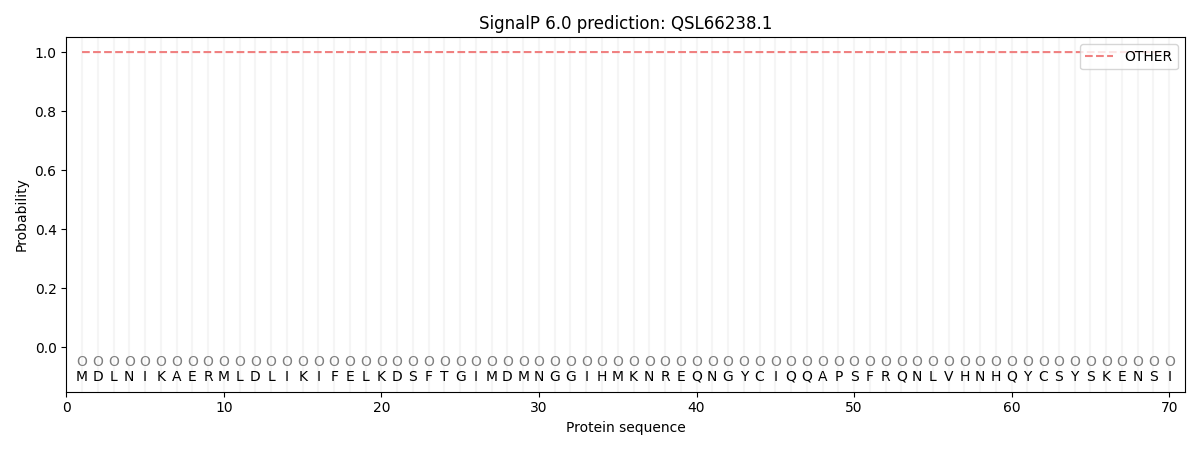
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000097 | 0.000000 |
