You are browsing environment: FUNGIDB
CAZyme Information: QRD05353.1
You are here: Home > Sequence: QRD05353.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Parastagonospora nodorum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Dothideomycetes; ; Phaeosphaeriaceae; Parastagonospora; Parastagonospora nodorum | |||||||||||
| CAZyme ID | QRD05353.1 | |||||||||||
| CAZy Family | GH92 | |||||||||||
| CAZyme Description | CBM1 domain-containing protein [Source:UniProtKB/TrEMBL;Acc:A0A7U2NP21] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 1.1.99.18:13 | 1.1.99.18:13 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA3 | 236 | 776 | 1.5e-242 | 0.9835766423357665 |
| AA8 | 27 | 208 | 6.2e-52 | 0.9945054945054945 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 406418 | CDH-cyt | 4.11e-58 | 27 | 204 | 1 | 177 | Cytochrome domain of cellobiose dehydrogenase. CDH-cyt is the cytochrome domain, at the N-terminus, of cellobiose dehydrogenase. CDH-cyt folds as a beta sandwich with the topology of the antibody Fab V(H) domain and binds iron. The haem iron is ligated by Met83 and His181 in UniProtKB:Q01738. |
| 225186 | BetA | 1.34e-40 | 244 | 777 | 7 | 537 | Choline dehydrogenase or related flavoprotein [Lipid transport and metabolism, General function prediction only]. |
| 187688 | CDH_like_cytochrome | 1.91e-32 | 27 | 195 | 2 | 166 | Heme-binding cytochrome domain of fungal cellobiose dehydrogenases. Cellobiose dehydrogenase (CellobioseDH or CDH) is an extracellular fungal oxidoreductase that degrades both lignin and cellulose. Specifically, CDHs oxidize cellobiose, cellodextrins, and lactose to corresponding lactones, utilizing a variety of electron acceptors. Class-II CDHs are monomeric hemoflavoenzymes that are comprised of a b-type cytochrome domain linked to a large flavodehydrogenase domain. The cytochrome domain of CDH and related enzymes, which this model describes, folds as a beta sandwich and complexes a heme molecule. It is found at the N-terminus of this family of enzymes, and belongs to the DOMON domain superfamily, a ligand-interacting motif found in all three kingdoms of life. |
| 235000 | PRK02106 | 3.19e-26 | 427 | 774 | 196 | 532 | choline dehydrogenase; Validated |
| 366272 | GMC_oxred_N | 1.63e-21 | 325 | 542 | 23 | 217 | GMC oxidoreductase. This family of proteins bind FAD as a cofactor. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 860 | 1 | 860 | |
| 0.0 | 1 | 859 | 1 | 859 | |
| 0.0 | 1 | 781 | 1 | 790 | |
| 0.0 | 1 | 858 | 1 | 837 | |
| 4.61e-298 | 1 | 858 | 1 | 830 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.11e-283 | 27 | 858 | 8 | 807 | Chain A, Cellobiose dehydrogenase [Thermothelomyces myriococcoides] |
|
| 1.02e-273 | 23 | 858 | 3 | 806 | Cellobiose dehydrogenase from Neurospora crassa, NcCDH [Neurospora crassa OR74A],4QI7_B Cellobiose dehydrogenase from Neurospora crassa, NcCDH [Neurospora crassa OR74A] |
|
| 3.92e-239 | 239 | 858 | 1 | 585 | Chain A, Cellobiose dehydrogenase [Thermothelomyces myriococcoides],4QI5_A Chain A, Cellobiose dehydrogenase [Thermothelomyces myriococcoides] |
|
| 3.76e-128 | 245 | 777 | 3 | 538 | Chain A, Cellobiose dehydrogenase [Phanerodontia chrysosporium],1NAA_B Chain B, Cellobiose dehydrogenase [Phanerodontia chrysosporium] |
|
| 4.38e-128 | 245 | 777 | 8 | 543 | Chain A, cellobiose dehydrogenase [Phanerodontia chrysosporium],1KDG_B Chain B, cellobiose dehydrogenase [Phanerodontia chrysosporium] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.35e-153 | 7 | 777 | 1 | 770 | Cellobiose dehydrogenase OS=Phanerodontia chrysosporium OX=2822231 GN=CDH-1 PE=1 SV=1 |
|
| 3.30e-22 | 242 | 770 | 2 | 524 | L-sorbose 1-dehydrogenase OS=Gluconobacter oxydans OX=442 PE=3 SV=1 |
|
| 6.60e-22 | 233 | 771 | 34 | 594 | Fatty acid photodecarboxylase, chloroplastic OS=Chlamydomonas reinhardtii OX=3055 GN=FAP PE=1 SV=1 |
|
| 1.31e-20 | 246 | 770 | 6 | 523 | Uncharacterized GMC-type oxidoreductase MT1316 OS=Mycobacterium tuberculosis (strain CDC 1551 / Oshkosh) OX=83331 GN=MT1316 PE=3 SV=1 |
|
| 1.31e-20 | 246 | 770 | 6 | 523 | Uncharacterized GMC-type oxidoreductase Mb1310 OS=Mycobacterium bovis (strain ATCC BAA-935 / AF2122/97) OX=233413 GN=BQ2027_MB1310 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
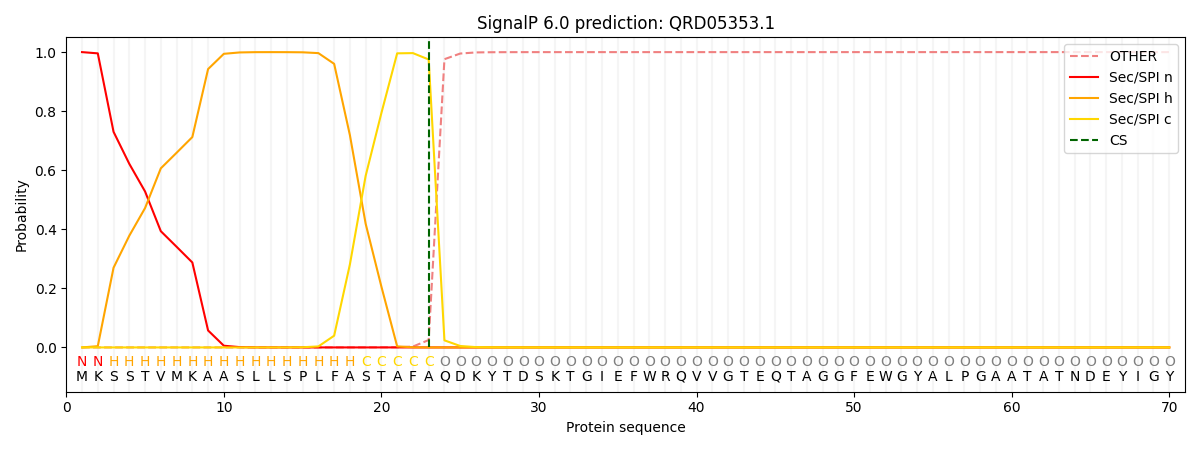
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000420 | 0.999538 | CS pos: 23-24. Pr: 0.9750 |
