You are browsing environment: FUNGIDB
CAZyme Information: QRC91755.1
You are here: Home > Sequence: QRC91755.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Parastagonospora nodorum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Dothideomycetes; ; Phaeosphaeriaceae; Parastagonospora; Parastagonospora nodorum | |||||||||||
| CAZyme ID | QRC91755.1 | |||||||||||
| CAZy Family | AA7 | |||||||||||
| CAZyme Description | Xyloglucan-specific endo-beta-1,4-glucanase A [Source:UniProtKB/TrEMBL;Acc:A0A7U2HTY7] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.4:15 | 3.2.1.151:9 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH12 | 131 | 275 | 1.8e-41 | 0.967948717948718 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396303 | Glyco_hydro_12 | 3.34e-58 | 72 | 276 | 1 | 205 | Glycosyl hydrolase family 12. |
| 235746 | PRK06215 | 1.06e-25 | 26 | 278 | 6 | 238 | hypothetical protein; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.65e-200 | 1 | 278 | 1 | 278 | |
| 2.10e-108 | 30 | 276 | 497 | 739 | |
| 3.27e-102 | 30 | 275 | 1 | 242 | |
| 1.04e-100 | 37 | 276 | 9 | 242 | |
| 1.04e-100 | 37 | 276 | 9 | 242 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.86e-86 | 55 | 276 | 7 | 227 | Crystal structure of XEG [Aspergillus aculeatus],3VL9_A Crystal structure of xeg-xyloglucan [Aspergillus aculeatus],3VL9_B Crystal structure of xeg-xyloglucan [Aspergillus aculeatus] |
|
| 2.40e-85 | 62 | 276 | 27 | 241 | Crystal Structure of the Family 12 Xyloglucanase from Aspergillus niveus [Aspergillus niveus],4NPR_B Crystal Structure of the Family 12 Xyloglucanase from Aspergillus niveus [Aspergillus niveus] |
|
| 1.96e-84 | 60 | 276 | 5 | 220 | Crystal structure of xeg-edgp [Aspergillus aculeatus],3VLB_D Crystal structure of xeg-edgp [Aspergillus aculeatus] |
|
| 1.01e-57 | 60 | 276 | 1 | 217 | Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 [Aspergillus aculeatus],5GM3_B Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 [Aspergillus aculeatus] |
|
| 5.91e-57 | 59 | 276 | 1 | 218 | Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellobiose [Aspergillus aculeatus],5GM5_B Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellobiose [Aspergillus aculeatus],5GM5_C Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellobiose [Aspergillus aculeatus],5GM5_D Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellobiose [Aspergillus aculeatus],5GM5_E Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellobiose [Aspergillus aculeatus],5GM5_F Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellobiose [Aspergillus aculeatus],5GM5_G Crystal structure of FI-CMCase from Aspergillus aculeatus F-50 in complex with cellobiose [Aspergillus aculeatus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.51e-99 | 30 | 276 | 1 | 248 | Probable xyloglucan-specific endo-beta-1,4-glucanase A OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=xgeA PE=3 SV=2 |
|
| 3.96e-97 | 30 | 278 | 1 | 239 | Xyloglucan-specific endo-beta-1,4-glucanase A OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=xgeA PE=1 SV=1 |
|
| 6.81e-91 | 56 | 276 | 18 | 238 | Probable xyloglucan-specific endo-beta-1,4-glucanase A OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=xgeA PE=3 SV=1 |
|
| 6.81e-91 | 56 | 276 | 18 | 238 | Probable xyloglucan-specific endo-beta-1,4-glucanase A OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=xgeA PE=3 SV=1 |
|
| 1.12e-86 | 30 | 276 | 1 | 236 | Xyloglucan-specific endo-beta-1,4-glucanase A OS=Aspergillus aculeatus OX=5053 GN=xgeA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
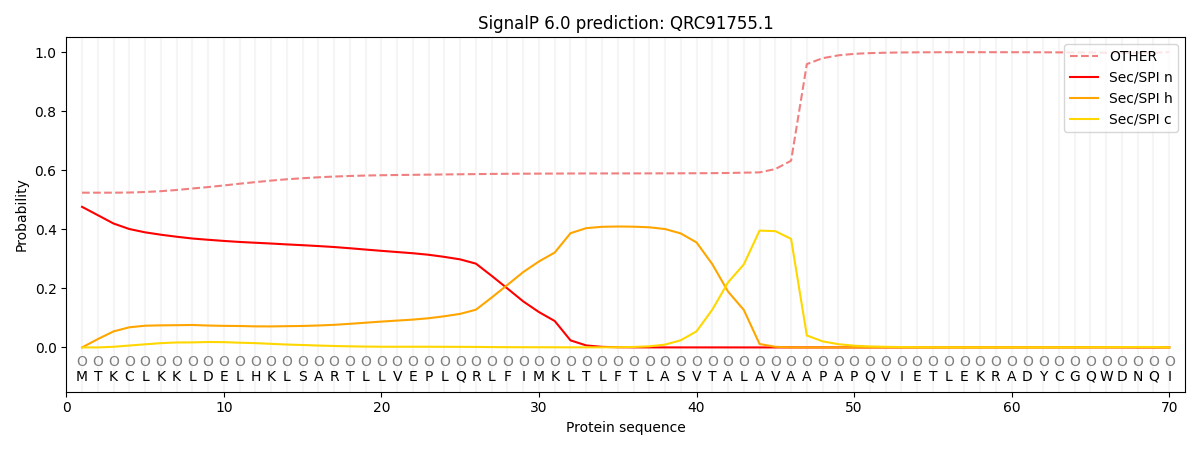
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.538319 | 0.461679 |

