You are browsing environment: FUNGIDB
CAZyme Information: QNG16379.1
You are here: Home > Sequence: QNG16379.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | [Candida] glabrata | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Saccharomycetes; ; Debaryomycetaceae; Candida; [Candida] glabrata | |||||||||||
| CAZyme ID | QNG16379.1 | |||||||||||
| CAZy Family | GT48 | |||||||||||
| CAZyme Description | SUN4 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH132 | 56 | 345 | 3.3e-120 | 0.966996699669967 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 397780 | SUN | 2.81e-139 | 92 | 332 | 1 | 244 | Beta-glucosidase (SUN family). Members of this family include Nca3, Sun4 and Sim1. This is a family of yeast proteins, involved in a diverse set of functions (DNA replication, aging, mitochondrial biogenesis and cell septation). BGLA from Candida wickerhamii has been characterized as a Beta-glucosidase EC:3.2.1.21. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.01e-246 | 1 | 345 | 1 | 345 | |
| 1.67e-245 | 1 | 345 | 1 | 345 | |
| 1.67e-245 | 1 | 345 | 1 | 345 | |
| 1.67e-245 | 1 | 345 | 1 | 345 | |
| 1.67e-245 | 1 | 345 | 1 | 345 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.03e-130 | 1 | 345 | 1 | 337 | Beta-glucosidase-like protein NCA3, mitochondrial OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=NCA3 PE=3 SV=1 |
|
| 6.81e-125 | 87 | 345 | 103 | 362 | Probable secreted beta-glucosidase UTH1 OS=Saccharomyces cerevisiae (strain RM11-1a) OX=285006 GN=UTH1 PE=3 SV=1 |
|
| 6.81e-125 | 87 | 345 | 103 | 362 | Probable secreted beta-glucosidase UTH1 OS=Saccharomyces cerevisiae (strain YJM789) OX=307796 GN=UTH1 PE=3 SV=2 |
|
| 6.81e-125 | 87 | 345 | 103 | 362 | Probable secreted beta-glucosidase UTH1 OS=Saccharomyces cerevisiae (strain AWRI1631) OX=545124 GN=UTH1 PE=3 SV=2 |
|
| 7.53e-125 | 87 | 345 | 106 | 365 | Probable secreted beta-glucosidase UTH1 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=UTH1 PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
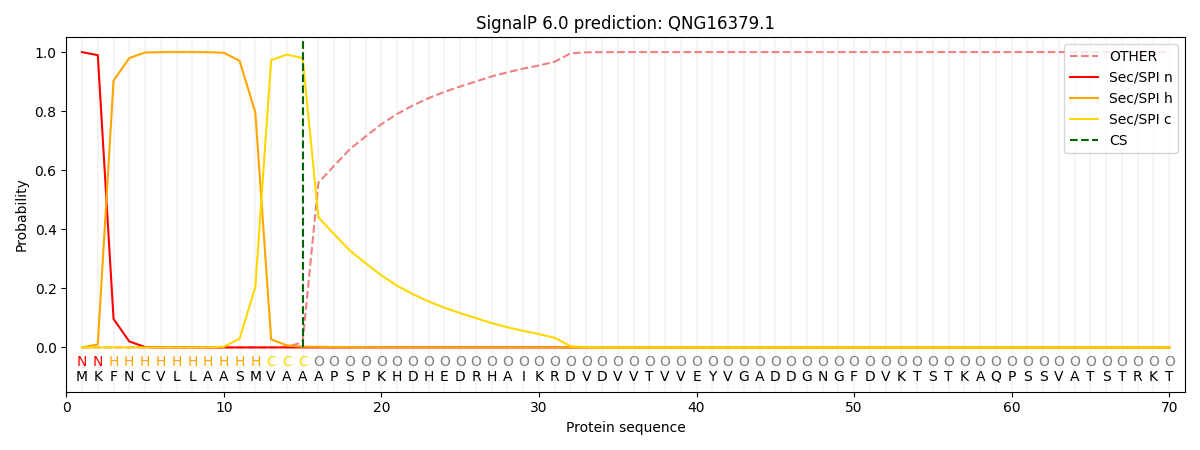
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000272 | 0.999692 | CS pos: 15-16. Pr: 0.9797 |
