You are browsing environment: FUNGIDB
CAZyme Information: QNG12164.1
You are here: Home > Sequence: QNG12164.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | [Candida] glabrata | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Saccharomycetes; ; Debaryomycetaceae; Candida; [Candida] glabrata | |||||||||||
| CAZyme ID | QNG12164.1 | |||||||||||
| CAZy Family | AA1 | |||||||||||
| CAZyme Description | CWH41 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.106:25 | 3.2.1.-:4 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH63 | 588 | 810 | 1.2e-31 | 0.43333333333333335 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 397353 | Glyco_hydro_63 | 0.0 | 309 | 812 | 1 | 493 | Glycosyl hydrolase family 63 C-terminal domain. This is a family of eukaryotic enzymes belonging to glycosyl hydrolase family 63. They catalyze the specific cleavage of the non-reducing terminal glucose residue from Glc(3)Man(9)GlcNAc(2). Mannosyl oligosaccharide glucosidase EC:3.2.1.106 is the first enzyme in the N-linked oligosaccharide processing pathway. This family represents the C-terminal catalytic domain. |
| 407154 | Glyco_hydro_63N | 3.17e-53 | 36 | 255 | 1 | 218 | Glycosyl hydrolase family 63 N-terminal domain. This is a family of eukaryotic enzymes belonging to glycosyl hydrolase family 63. They catalyze the specific cleavage of the non-reducing terminal glucose residue from Glc(3)Man(9)GlcNAc(2). Mannosyl oligosaccharide glucosidase EC:3.2.1.106 is the first enzyme in the N-linked oligosaccharide processing pathway. This family represents the N-terminal beta sandwich domain. |
| 225942 | GDB1 | 1.37e-09 | 360 | 742 | 245 | 556 | Glycogen debranching enzyme (alpha-1,6-glucosidase) [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 814 | 1 | 814 | |
| 0.0 | 1 | 814 | 1 | 814 | |
| 0.0 | 1 | 814 | 1 | 814 | |
| 0.0 | 1 | 814 | 1 | 814 | |
| 0.0 | 1 | 814 | 1 | 814 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.11e-305 | 23 | 811 | 2 | 800 | Crystal structure of Processing alpha-Glucosidase I [Saccharomyces cerevisiae S288C] |
|
| 1.48e-87 | 36 | 812 | 37 | 776 | Murine endoplasmic reticulum alpha-glucosidase I with N-9'-methoxynonyl-1-deoxynojirimycin [Mus musculus],5MHF_B Murine endoplasmic reticulum alpha-glucosidase I with N-9'-methoxynonyl-1-deoxynojirimycin [Mus musculus],5MHF_C Murine endoplasmic reticulum alpha-glucosidase I with N-9'-methoxynonyl-1-deoxynojirimycin [Mus musculus],5MHF_D Murine endoplasmic reticulum alpha-glucosidase I with N-9'-methoxynonyl-1-deoxynojirimycin [Mus musculus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.06e-305 | 17 | 811 | 26 | 830 | Mannosyl-oligosaccharide glucosidase OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=CWH41 PE=1 SV=1 |
|
| 5.14e-163 | 26 | 814 | 32 | 807 | Probable mannosyl-oligosaccharide glucosidase OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=SPAC6G10.09 PE=3 SV=1 |
|
| 9.30e-91 | 36 | 808 | 94 | 830 | Mannosyl-oligosaccharide glucosidase OS=Homo sapiens OX=9606 GN=MOGS PE=1 SV=5 |
|
| 4.49e-89 | 36 | 812 | 92 | 831 | Mannosyl-oligosaccharide glucosidase OS=Mus musculus OX=10090 GN=Mogs PE=1 SV=1 |
|
| 6.24e-88 | 28 | 812 | 101 | 846 | Mannosyl-oligosaccharide glucosidase GCS1 OS=Arabidopsis thaliana OX=3702 GN=GCS1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
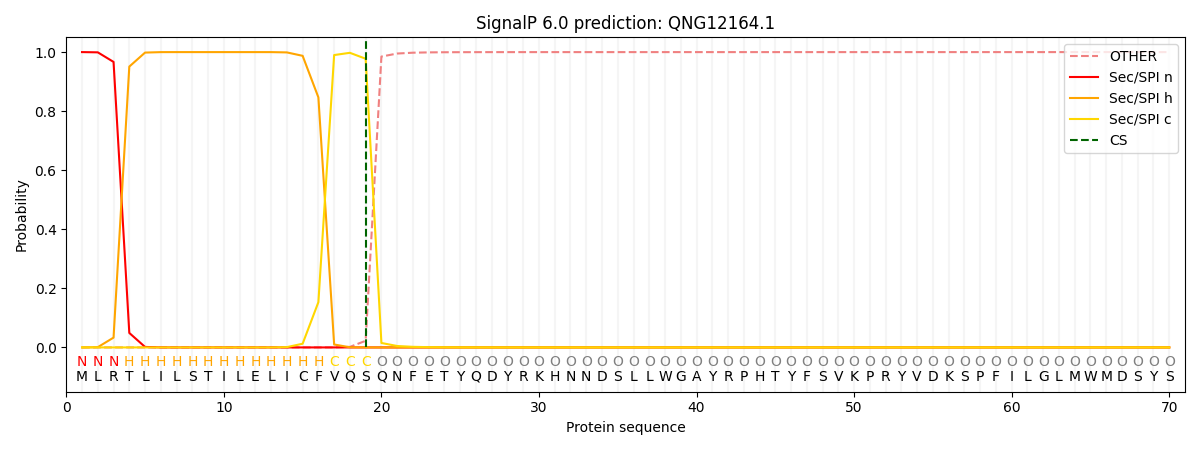
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000266 | 0.999722 | CS pos: 19-20. Pr: 0.9775 |
