You are browsing environment: FUNGIDB
CAZyme Information: QHS75578.1
You are here: Home > Sequence: QHS75578.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
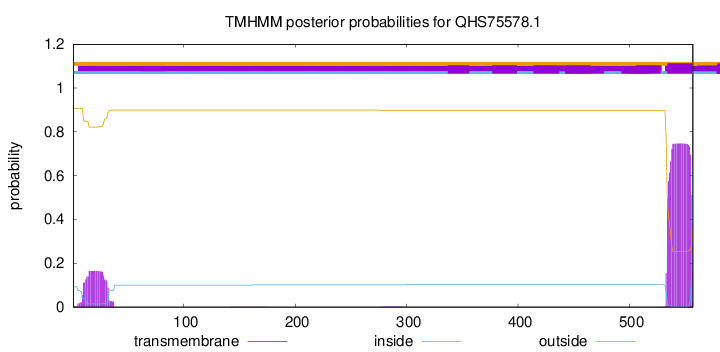
TMHMM annotations
Basic Information help
| Species | Saccharomyces paradoxus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Saccharomycetes; ; Saccharomycetaceae; Saccharomyces; Saccharomyces paradoxus | |||||||||||
| CAZyme ID | QHS75578.1 | |||||||||||
| CAZy Family | GT33 | |||||||||||
| CAZyme Description | Gas1 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 2.4.1.-:30 | 2.4.1.-:33 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH72 | 22 | 329 | 1.2e-140 | 0.9935897435897436 |
| CBM43 | 378 | 462 | 4.3e-23 | 0.9759036144578314 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 397351 | Glyco_hydro_72 | 0.0 | 17 | 330 | 1 | 315 | Glucanosyltransferase. This is a family of glycosylphosphatidylinositol-anchored beta(1-3)glucanosyltransferases. The active site residues in the Aspergillus fumigatus example are the two glutamate residues at 160 and 261. |
| 400371 | X8 | 4.91e-27 | 390 | 452 | 14 | 76 | X8 domain. The X8 domain domain contains at least 6 conserved cysteine residues that presumably form three disulphide bridges. The domain is found in an Olive pollen allergen as well as at the C-terminus of several families of glycosyl hydrolases. This domain may be involved in carbohydrate binding. This domain is characteristic of GPI-anchored domains. |
| 197867 | X8 | 1.27e-17 | 390 | 464 | 14 | 85 | Possibly involved in carbohydrate binding. The X8 domain, which may be involved in carbohydrate binding, is found in an Olive pollen antigen as well as at the C terminus of family 17 glycosyl hydrolases. It contains 6 conserved cysteine residues which presumably form three disulfide bridges. |
| 225789 | LacZ | 2.47e-05 | 28 | 237 | 286 | 474 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 557 | 1 | 557 | |
| 0.0 | 1 | 557 | 1 | 558 | |
| 0.0 | 1 | 557 | 1 | 558 | |
| 0.0 | 1 | 557 | 1 | 558 | |
| 0.0 | 1 | 557 | 1 | 558 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.12e-141 | 26 | 445 | 34 | 466 | Saccharomyces cerevisiae Gas2p in complex with laminaripentaose [Saccharomyces cerevisiae],2W63_A Saccharomyces Cerevisiae Gas2p In Complex With Laminaritriose And Laminaritetraose [Saccharomyces cerevisiae],5O9O_A Crystal structure of ScGas2 in complex with compound 7. [Saccharomyces cerevisiae S288C],5O9P_A Crystal structure of Gas2 in complex with compound 10 [Saccharomyces cerevisiae S288C],5O9Q_A Crystal structure of ScGas2 in complex with compound 6 [Saccharomyces cerevisiae S288C],5O9R_A Crystal structure of ScGas2 in complex with compound 9 [Saccharomyces cerevisiae S288C],5O9Y_A Crystal structure of ScGas2 in complex with compound 11 [Saccharomyces cerevisiae S288C],5OA2_A Crystal structure of ScGas2 in complex with compound 8 [Saccharomyces cerevisiae S288C],5OA2_B Crystal structure of ScGas2 in complex with compound 8 [Saccharomyces cerevisiae S288C],5OA2_C Crystal structure of ScGas2 in complex with compound 8 [Saccharomyces cerevisiae S288C],5OA6_A Crystal structure of ScGas2 in complex with compound 12 [Saccharomyces cerevisiae S288C] |
|
| 5.98e-141 | 26 | 445 | 34 | 466 | Saccharomyces cerevisiae Gas2p apostructure (E176Q mutant) [Saccharomyces cerevisiae] |
|
| 5.98e-141 | 26 | 445 | 34 | 466 | SACCHAROMYCES CEREVISIAE GAS2P (E176Q MUTANT) IN COMPLEX WITH LAMINARITETRAOSE AND LAMINARIPENTAOSE [Saccharomyces cerevisiae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 0.0 | 1 | 557 | 1 | 559 | 1,3-beta-glucanosyltransferase GAS1 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=GAS1 PE=1 SV=2 |
|
| 1.49e-207 | 1 | 481 | 1 | 480 | Protein EPD1 OS=Candida maltosa OX=5479 GN=EPD1 PE=3 SV=1 |
|
| 4.41e-203 | 1 | 481 | 1 | 480 | pH-responsive protein 2 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=PHR2 PE=2 SV=2 |
|
| 2.01e-187 | 26 | 480 | 27 | 488 | pH-responsive protein 1 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=PHR1 PE=2 SV=4 |
|
| 7.19e-183 | 5 | 465 | 10 | 477 | Protein EPD2 OS=Candida maltosa OX=5479 GN=EPD2 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000242 | 0.999720 | CS pos: 22-23. Pr: 0.9805 |