You are browsing environment: FUNGIDB
CAZyme Information: QHS73583.1
You are here: Home > Sequence: QHS73583.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Saccharomyces paradoxus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Saccharomycetes; ; Saccharomycetaceae; Saccharomyces; Saccharomyces paradoxus | |||||||||||
| CAZyme ID | QHS73583.1 | |||||||||||
| CAZy Family | GH47 | |||||||||||
| CAZyme Description | Ima1 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.10:2 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH13 | 37 | 399 | 2.2e-186 | 0.9972144846796658 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 200472 | AmyAc_SI_OligoGlu_DGase | 0.0 | 16 | 503 | 1 | 427 | Alpha amylase catalytic domain found in Sucrose isomerases, oligo-1,6-glucosidase (also called isomaltase; sucrase-isomaltase; alpha-limit dextrinase), dextran glucosidase (also called glucan 1,6-alpha-glucosidase), and related proteins. The sucrose isomerases (SIs) Isomaltulose synthase (EC 5.4.99.11) and Trehalose synthase (EC 5.4.99.16) catalyze the isomerization of sucrose and maltose to produce isomaltulose and trehalulose, respectively. Oligo-1,6-glucosidase (EC 3.2.1.10) hydrolyzes the alpha-1,6-glucosidic linkage of isomaltooligosaccharides, pannose, and dextran. Unlike alpha-1,4-glucosidases (EC 3.2.1.20), it fails to hydrolyze the alpha-1,4-glucosidic bonds of maltosaccharides. Dextran glucosidase (DGase, EC 3.2.1.70) hydrolyzes alpha-1,6-glucosidic linkages at the non-reducing end of panose, isomaltooligosaccharides and dextran to produce alpha-glucose.The common reaction chemistry of the alpha-amylase family enzymes is based on a two-step acid catalytic mechanism that requires two critical carboxylates: one acting as a general acid/base (Glu) and the other as a nucleophile (Asp). Both hydrolysis and transglycosylation proceed via the nucleophilic substitution reaction between the anomeric carbon, C1 and a nucleophile. Both enzymes contain the three catalytic residues (Asp, Glu and Asp) common to the alpha-amylase family as well as two histidine residues which are predicted to be critical to binding the glucose residue adjacent to the scissile bond in the substrates. The Alpha-amylase family comprises the largest family of glycoside hydrolases (GH), with the majority of enzymes acting on starch, glycogen, and related oligo- and polysaccharides. These proteins catalyze the transformation of alpha-1,4 and alpha-1,6 glucosidic linkages with retention of the anomeric center. The protein is described as having 3 domains: A, B, C. A is a (beta/alpha) 8-barrel; B is a loop between the beta 3 strand and alpha 3 helix of A; C is the C-terminal extension characterized by a Greek key. The majority of the enzymes have an active site cleft found between domains A and B where a triad of catalytic residues performs catalysis. Other members of this family have lost the catalytic activity as in the case of the human 4F2hc, or only have 2 residues that serve as the catalytic nucleophile and the acid/base, such as Thermus A4 beta-galactosidase with 2 Glu residues (GH42) and human alpha-galactosidase with 2 Asp residues (GH31). The family members are quite extensive and include: alpha amylase, maltosyltransferase, cyclodextrin glycotransferase, maltogenic amylase, neopullulanase, isoamylase, 1,4-alpha-D-glucan maltotetrahydrolase, 4-alpha-glucotransferase, oligo-1,6-glucosidase, amylosucrase, sucrose phosphorylase, and amylomaltase. |
| 274115 | trehalose_treC | 8.54e-174 | 14 | 587 | 1 | 542 | alpha,alpha-phosphotrehalase. Trehalose is a glucose disaccharide that serves in many biological systems as a compatible solute for protection against hyperosmotic and thermal stress. This family describes trehalose-6-phosphate hydrolase, product of the treC (or treA) gene, which is often found together with a trehalose uptake transporter and a trehalose operon repressor. |
| 395077 | Alpha-amylase | 8.10e-164 | 37 | 400 | 1 | 332 | Alpha amylase, catalytic domain. Alpha amylase is classified as family 13 of the glycosyl hydrolases. The structure is an 8 stranded alpha/beta barrel containing the active site, interrupted by a ~70 a.a. calcium-binding domain protruding between beta strand 3 and alpha helix 3, and a carboxyl-terminal Greek key beta-barrel domain. |
| 223443 | AmyA | 1.47e-145 | 18 | 555 | 1 | 495 | Glycosidase [Carbohydrate transport and metabolism]. |
| 182849 | PRK10933 | 2.66e-142 | 10 | 589 | 3 | 551 | trehalose-6-phosphate hydrolase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 589 | 1 | 589 | |
| 0.0 | 1 | 589 | 1 | 589 | |
| 0.0 | 1 | 589 | 1 | 589 | |
| 0.0 | 1 | 589 | 1 | 589 | |
| 0.0 | 1 | 589 | 1 | 589 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 0.0 | 1 | 589 | 1 | 589 | Crystal structure of isomaltase in complex with isomaltose [Saccharomyces cerevisiae],3AXI_A Crystal structure of isomaltase in complex with maltose [Saccharomyces cerevisiae] |
|
| 0.0 | 1 | 589 | 1 | 589 | Crystal structure of isomaltase from Saccharomyces cerevisiae [Saccharomyces cerevisiae],3A4A_A Crystal structure of isomaltase from Saccharomyces cerevisiae [Saccharomyces cerevisiae],3AJ7_A Crystal Structure of isomaltase from Saccharomyces cerevisiae [Saccharomyces cerevisiae] |
|
| 2.07e-174 | 9 | 586 | 11 | 578 | Chain A, BaAG2 [Blastobotrys adeninivorans],7P01_B Chain B, BaAG2 [Blastobotrys adeninivorans],7P07_A Chain A, BaAG2 [Blastobotrys adeninivorans],7P07_B Chain B, BaAG2 [Blastobotrys adeninivorans] |
|
| 6.24e-162 | 13 | 588 | 3 | 560 | The Structure of Wild-type MalL from Bacillus subtilis [Bacillus subtilis subsp. subtilis str. 168],4M56_B The Structure of Wild-type MalL from Bacillus subtilis [Bacillus subtilis subsp. subtilis str. 168] |
|
| 8.83e-162 | 13 | 588 | 3 | 560 | The Structure of MalL mutant enzyme G202P from Bacillus subtilus [Bacillus subtilis subsp. subtilis str. 168] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 0.0 | 1 | 589 | 1 | 589 | Oligo-1,6-glucosidase IMA4 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=IMA4 PE=3 SV=1 |
|
| 0.0 | 1 | 587 | 1 | 582 | Alpha-glucosidase MAL32 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=MAL32 PE=1 SV=1 |
|
| 0.0 | 1 | 589 | 1 | 589 | Oligo-1,6-glucosidase IMA1 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=IMA1 PE=1 SV=1 |
|
| 0.0 | 1 | 587 | 1 | 582 | Alpha-glucosidase MAL12 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=MAL12 PE=1 SV=1 |
|
| 0.0 | 1 | 589 | 1 | 589 | Oligo-1,6-glucosidase IMA2 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=IMA2 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
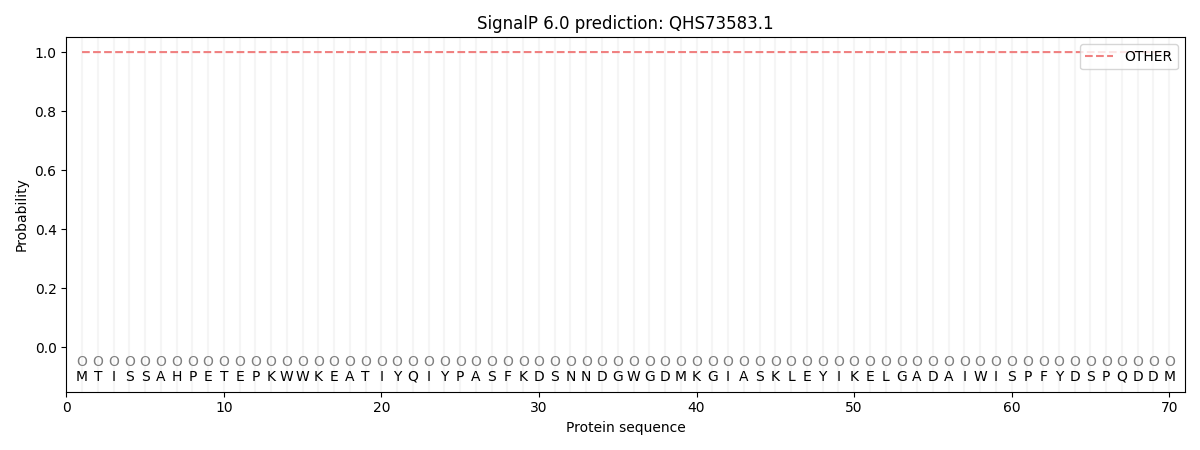
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000072 | 0.000005 |
