You are browsing environment: FUNGIDB
CAZyme Information: PYU1_T013516-p1
You are here: Home > Sequence: PYU1_T013516-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Globisporangium ultimum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Pythiaceae; Globisporangium; Globisporangium ultimum | |||||||||||
| CAZyme ID | PYU1_T013516-p1 | |||||||||||
| CAZy Family | GT71 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT30 | 102 | 226 | 1.3e-33 | 0.672316384180791 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 398219 | Glycos_transf_N | 2.95e-47 | 40 | 226 | 2 | 155 | 3-Deoxy-D-manno-octulosonic-acid transferase (kdotransferase). Members of this family transfer activated sugars to a variety of substrates, including glycogen, fructose-6-phosphate and lipopolysaccharides. Members of the family transfer UDP, ADP, GDP or CMP linked sugars. The Glycos_transf_N region is flanked at the N-terminus by a signal peptide and at the C-terminus by Glycos_transf_1 (pfam00534). The eukaryotic glycogen synthases may be distant members of this bacterial family. |
| 224436 | KdtA | 1.12e-43 | 7 | 221 | 3 | 186 | 3-deoxy-D-manno-octulosonic-acid transferase [Cell wall/membrane/envelope biogenesis]. |
| 235589 | PRK05749 | 1.36e-42 | 7 | 222 | 4 | 188 | 3-deoxy-D-manno-octulosonic-acid transferase; Reviewed |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.63e-33 | 4 | 217 | 1 | 182 | |
| 2.31e-32 | 4 | 217 | 1 | 182 | |
| 2.12e-29 | 5 | 221 | 1 | 185 | |
| 6.23e-29 | 8 | 220 | 12 | 192 | |
| 7.71e-28 | 9 | 222 | 6 | 191 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.74e-07 | 48 | 172 | 32 | 127 | Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, substrate-free form [Aquifex aeolicus],2XCI_B Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, substrate-free form [Aquifex aeolicus],2XCI_C Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, substrate-free form [Aquifex aeolicus],2XCI_D Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, substrate-free form [Aquifex aeolicus],2XCU_A Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, complex with CMP [Aquifex aeolicus],2XCU_B Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, complex with CMP [Aquifex aeolicus],2XCU_C Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, complex with CMP [Aquifex aeolicus],2XCU_D Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, complex with CMP [Aquifex aeolicus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.34e-20 | 1 | 216 | 3 | 188 | Probable 3-deoxy-D-manno-octulosonic acid transferase, mitochondrial OS=Arabidopsis thaliana OX=3702 GN=KDTA PE=2 SV=1 |
|
| 2.63e-19 | 24 | 222 | 26 | 185 | 3-deoxy-D-manno-octulosonic acid transferase OS=Rickettsia bellii (strain RML369-C) OX=336407 GN=waaA PE=3 SV=1 |
|
| 9.71e-18 | 45 | 222 | 85 | 232 | 3-deoxy-D-manno-octulosonic acid transferase OS=Rickettsia felis (strain ATCC VR-1525 / URRWXCal2) OX=315456 GN=waaA PE=3 SV=1 |
|
| 1.13e-16 | 62 | 222 | 93 | 227 | 3-deoxy-D-manno-octulosonic acid transferase OS=Rickettsia prowazekii (strain Madrid E) OX=272947 GN=waaA PE=3 SV=1 |
|
| 3.27e-15 | 62 | 221 | 95 | 228 | 3-deoxy-D-manno-octulosonic acid transferase OS=Rickettsia typhi (strain ATCC VR-144 / Wilmington) OX=257363 GN=waaA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
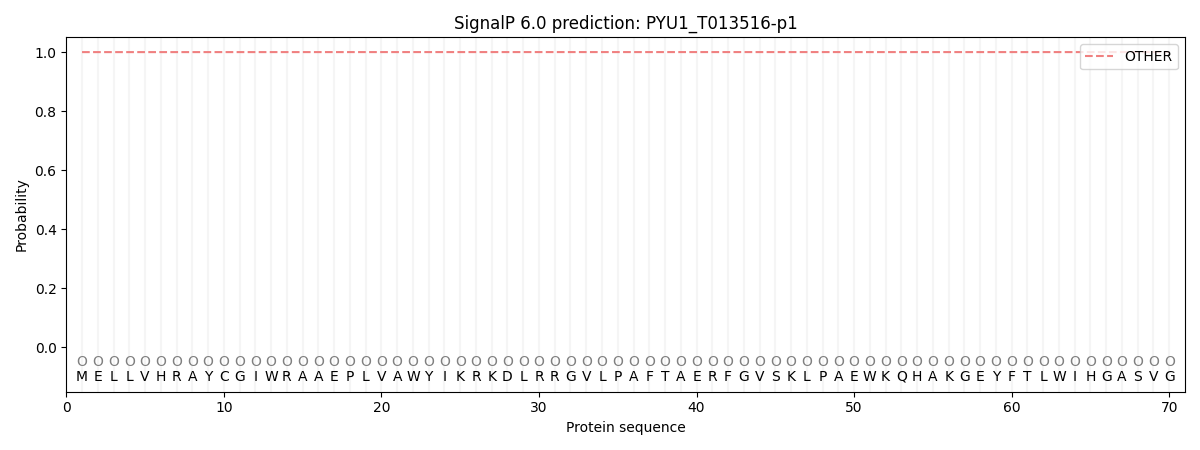
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000044 | 0.000000 |
