You are browsing environment: FUNGIDB
CAZyme Information: PYU1_T013487-p1
You are here: Home > Sequence: PYU1_T013487-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
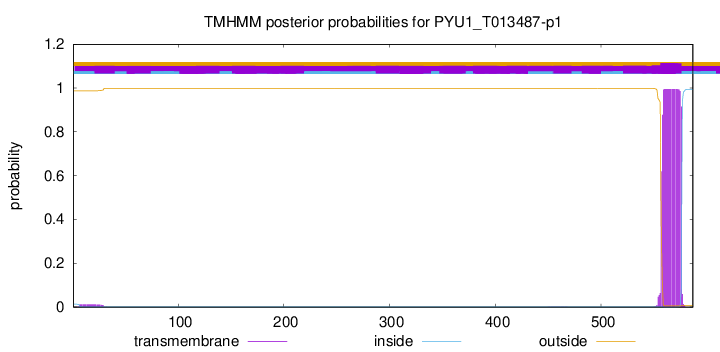
TMHMM annotations
Basic Information help
| Species | Globisporangium ultimum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Pythiaceae; Globisporangium; Globisporangium ultimum | |||||||||||
| CAZyme ID | PYU1_T013487-p1 | |||||||||||
| CAZy Family | GT71 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.21:2 | 3.2.1.23:1 | 3.2.1.38:1 | 3.2.1.-:1 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH1 | 37 | 521 | 2.4e-138 | 0.9906759906759907 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395176 | Glyco_hydro_1 | 6.31e-148 | 40 | 521 | 5 | 451 | Glycosyl hydrolase family 1. |
| 274539 | BGL | 1.16e-128 | 41 | 514 | 1 | 426 | beta-galactosidase. |
| 225343 | BglB | 8.73e-121 | 40 | 524 | 4 | 456 | Beta-glucosidase/6-phospho-beta-glucosidase/beta-galactosidase [Carbohydrate transport and metabolism]. |
| 215435 | PLN02814 | 1.73e-95 | 24 | 514 | 12 | 476 | beta-glucosidase |
| 215455 | PLN02849 | 2.98e-89 | 40 | 514 | 30 | 476 | beta-glucosidase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 2.09e-246 | 8 | 532 | 7 | 523 | |
| 9.23e-237 | 39 | 523 | 12 | 496 | |
| 2.47e-222 | 26 | 524 | 9 | 507 | |
| 4.15e-157 | 88 | 526 | 5 | 421 | |
| 5.90e-107 | 40 | 521 | 4 | 456 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.46e-95 | 26 | 516 | 25 | 503 | Crystal structure of beta-primeverosidase [Camellia sinensis],3WQ4_B Crystal structure of beta-primeverosidase [Camellia sinensis],3WQ5_A beta-Primeverosidase in complex with disaccharide substrate-analog N-beta-primeverosylamidine, natural aglycone derivative [Camellia sinensis],3WQ5_B beta-Primeverosidase in complex with disaccharide substrate-analog N-beta-primeverosylamidine, natural aglycone derivative [Camellia sinensis],3WQ6_A beta-Primeverosidase in complex with disaccharide substrate-analog N-beta-primeverosylamidine, artificial aglycone derivative [Camellia sinensis],3WQ6_B beta-Primeverosidase in complex with disaccharide substrate-analog N-beta-primeverosylamidine, artificial aglycone derivative [Camellia sinensis] |
|
| 1.17e-91 | 40 | 513 | 22 | 506 | Crystal structure of native Raucaffricine glucosidase [Rauvolfia serpentina],4ATD_B Crystal structure of native Raucaffricine glucosidase [Rauvolfia serpentina],4ATL_A Crystal structure of Raucaffricine glucosidase in complex with Glucose [Rauvolfia serpentina],4ATL_B Crystal structure of Raucaffricine glucosidase in complex with Glucose [Rauvolfia serpentina] |
|
| 1.25e-91 | 40 | 524 | 13 | 466 | Chimeric Family 1 beta-glucosidase made with non-contiguous SCHEMA [Thermotoga maritima MSB8],4GXP_B Chimeric Family 1 beta-glucosidase made with non-contiguous SCHEMA [Thermotoga maritima MSB8],4GXP_C Chimeric Family 1 beta-glucosidase made with non-contiguous SCHEMA [Thermotoga maritima MSB8] |
|
| 2.43e-91 | 40 | 513 | 22 | 506 | Crystal of Raucaffricine Glucosidase in complex with inhibitor [Rauvolfia serpentina],3ZJ6_B Crystal of Raucaffricine Glucosidase in complex with inhibitor [Rauvolfia serpentina],4A3Y_A Crystal structure of Raucaffricine glucosidase from ajmaline biosynthesis pathway [Rauvolfia serpentina],4A3Y_B Crystal structure of Raucaffricine glucosidase from ajmaline biosynthesis pathway [Rauvolfia serpentina] |
|
| 3.24e-91 | 40 | 513 | 22 | 506 | Structures of Alkaloid Biosynthetic Glucosidases Decode Substrate Specificity [Rauvolfia serpentina],3U57_B Structures of Alkaloid Biosynthetic Glucosidases Decode Substrate Specificity [Rauvolfia serpentina],3U5U_A Structures of Alkaloid Biosynthetic Glucosidases Decode Substrate Specificity [Rauvolfia serpentina],3U5U_B Structures of Alkaloid Biosynthetic Glucosidases Decode Substrate Specificity [Rauvolfia serpentina],3U5Y_A Structures of Alkaloid Biosynthetic Glucosidases Decode Substrate Specificity [Rauvolfia serpentina],3U5Y_B Structures of Alkaloid Biosynthetic Glucosidases Decode Substrate Specificity [Rauvolfia serpentina],4EK7_A High speed X-ray analysis of plant enzymes at room temperature [Rauvolfia serpentina],4EK7_B High speed X-ray analysis of plant enzymes at room temperature [Rauvolfia serpentina] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.06e-95 | 36 | 515 | 37 | 507 | Beta-glucosidase 21 OS=Arabidopsis thaliana OX=3702 GN=BGLU21 PE=1 SV=1 |
|
| 2.14e-94 | 40 | 515 | 37 | 506 | Beta-glucosidase 32 OS=Arabidopsis thaliana OX=3702 GN=BGLU32 PE=2 SV=2 |
|
| 4.52e-94 | 24 | 515 | 15 | 507 | Beta-glucosidase 22 OS=Arabidopsis thaliana OX=3702 GN=BGLU22 PE=1 SV=1 |
|
| 9.28e-93 | 40 | 516 | 19 | 490 | Beta-glucosidase 26, peroxisomal OS=Arabidopsis thaliana OX=3702 GN=BGLU26 PE=1 SV=1 |
|
| 2.52e-92 | 40 | 519 | 37 | 510 | Beta-glucosidase 31 OS=Arabidopsis thaliana OX=3702 GN=BGLU31 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000420 | 0.999551 | CS pos: 20-21. Pr: 0.9723 |