You are browsing environment: FUNGIDB
CAZyme Information: PWY92018.1
You are here: Home > Sequence: PWY92018.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus heteromorphus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus heteromorphus | |||||||||||
| CAZyme ID | PWY92018.1 | |||||||||||
| CAZy Family | GT35 | |||||||||||
| CAZyme Description | putative beta-glucosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.21:13 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH1 | 8 | 477 | 8e-163 | 0.9836829836829837 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 274539 | BGL | 0.0 | 11 | 470 | 1 | 424 | beta-galactosidase. |
| 395176 | Glyco_hydro_1 | 0.0 | 9 | 475 | 4 | 447 | Glycosyl hydrolase family 1. |
| 225343 | BglB | 5.48e-146 | 7 | 477 | 1 | 451 | Beta-glucosidase/6-phospho-beta-glucosidase/beta-galactosidase [Carbohydrate transport and metabolism]. |
| 215455 | PLN02849 | 1.91e-107 | 3 | 469 | 23 | 473 | beta-glucosidase |
| 215435 | PLN02814 | 1.99e-106 | 6 | 469 | 24 | 473 | beta-glucosidase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 482 | 1 | 482 | |
| 0.0 | 1 | 482 | 1 | 482 | |
| 0.0 | 1 | 482 | 1 | 482 | |
| 0.0 | 1 | 482 | 1 | 482 | |
| 0.0 | 1 | 482 | 1 | 482 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.84e-270 | 2 | 482 | 18 | 498 | Chain A, Beta-glucosidase [[Humicola] grisea var. thermoidea],4MDP_A Chain A, Beta-glucosidase [[Humicola] grisea var. thermoidea] |
|
| 9.83e-244 | 10 | 482 | 3 | 465 | Crystal structure of the double mutant L167W / P172L of the beta-glucosidase from Trichoderma harzianum [Trichoderma harzianum],6EFU_B Crystal structure of the double mutant L167W / P172L of the beta-glucosidase from Trichoderma harzianum [Trichoderma harzianum] |
|
| 1.08e-240 | 10 | 482 | 3 | 465 | Trichoderma harzianum GH1 beta-glucosidase ThBgl1 [Trichoderma harzianum],5JBK_B Trichoderma harzianum GH1 beta-glucosidase ThBgl1 [Trichoderma harzianum] |
|
| 2.17e-240 | 10 | 482 | 22 | 484 | Crystal structure of the beta-glucosidase from Trichoderma harzianum [Trichoderma harzianum],5BWF_B Crystal structure of the beta-glucosidase from Trichoderma harzianum [Trichoderma harzianum] |
|
| 2.78e-233 | 10 | 480 | 9 | 470 | Chain A, Beta-glucosidase [Trichoderma reesei],3AHY_B Chain B, Beta-glucosidase [Trichoderma reesei],3AHY_C Chain C, Beta-glucosidase [Trichoderma reesei],3AHY_D Chain D, Beta-glucosidase [Trichoderma reesei] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.31e-191 | 10 | 480 | 11 | 475 | Beta-glucosidase 1B OS=Phanerodontia chrysosporium OX=2822231 GN=BGL1B PE=1 SV=1 |
|
| 9.11e-173 | 10 | 481 | 6 | 461 | Beta-glucosidase 1A OS=Phanerodontia chrysosporium OX=2822231 GN=BGL1A PE=1 SV=1 |
|
| 5.14e-144 | 2 | 473 | 25 | 499 | Beta-glucosidase 24 OS=Oryza sativa subsp. japonica OX=39947 GN=BGLU24 PE=2 SV=1 |
|
| 5.02e-143 | 9 | 469 | 38 | 501 | Beta-glucosidase 12 OS=Oryza sativa subsp. japonica OX=39947 GN=BGLU12 PE=1 SV=2 |
|
| 2.01e-142 | 9 | 469 | 38 | 501 | Beta-glucosidase 12 OS=Oryza sativa subsp. indica OX=39946 GN=BGLU12 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
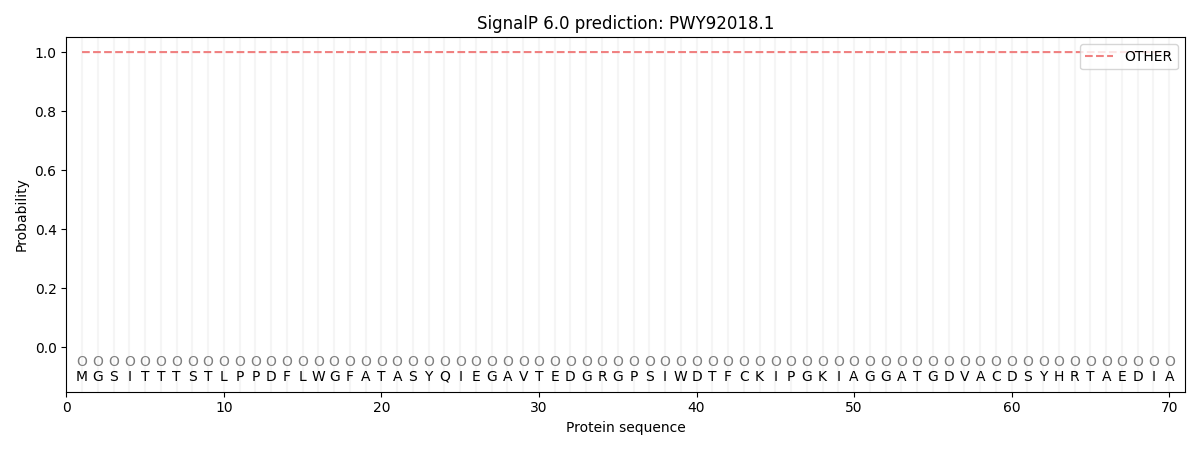
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000071 | 0.000001 |
