You are browsing environment: FUNGIDB
CAZyme Information: PWY83012.1
You are here: Home > Sequence: PWY83012.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus heteromorphus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus heteromorphus | |||||||||||
| CAZyme ID | PWY83012.1 | |||||||||||
| CAZy Family | GH35 | |||||||||||
| CAZyme Description | MIPC synthase subunit | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT32 | 75 | 155 | 1.1e-23 | 0.9444444444444444 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 226297 | OCH1 | 2.68e-80 | 41 | 286 | 63 | 311 | Mannosyltransferase OCH1 or related enzyme [Cell wall/membrane/envelope biogenesis]. |
| 398274 | Gly_transf_sug | 3.17e-27 | 73 | 157 | 1 | 89 | Glycosyltransferase sugar-binding region containing DXD motif. The DXD motif is a short conserved motif found in many families of glycosyltransferases, which add a range of different sugars to other sugars, phosphates and proteins. DXD-containing glycosyltransferases all use nucleoside diphosphate sugars as donors and require divalent cations, usually manganese. The DXD motif is expected to play a carbohydrate binding role in sugar-nucleoside diphosphate and manganese dependent glycosyltransferases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.51e-257 | 1 | 359 | 1 | 359 | |
| 4.51e-257 | 1 | 359 | 1 | 359 | |
| 9.09e-257 | 1 | 359 | 1 | 359 | |
| 3.72e-237 | 1 | 359 | 1 | 356 | |
| 5.69e-237 | 1 | 359 | 1 | 358 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.13e-106 | 12 | 279 | 12 | 275 | Mannosyl phosphorylinositol ceramide synthase SUR1 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=SUR1 PE=1 SV=1 |
|
| 1.68e-104 | 1 | 279 | 1 | 283 | Mannosyl phosphorylinositol ceramide synthase CSH1 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=CSH1 PE=1 SV=1 |
|
| 4.77e-91 | 37 | 286 | 56 | 312 | Inositol phosphoceramide mannosyltransferase 2 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=SPCC4F11.04c PE=3 SV=1 |
|
| 1.56e-86 | 48 | 284 | 46 | 283 | Inositol phosphoceramide mannosyltransferase 3 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=SPAC17G8.11c PE=1 SV=1 |
|
| 1.56e-54 | 59 | 269 | 64 | 275 | Inositol phosphoceramide mannosyltransferase 1 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=imt1 PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
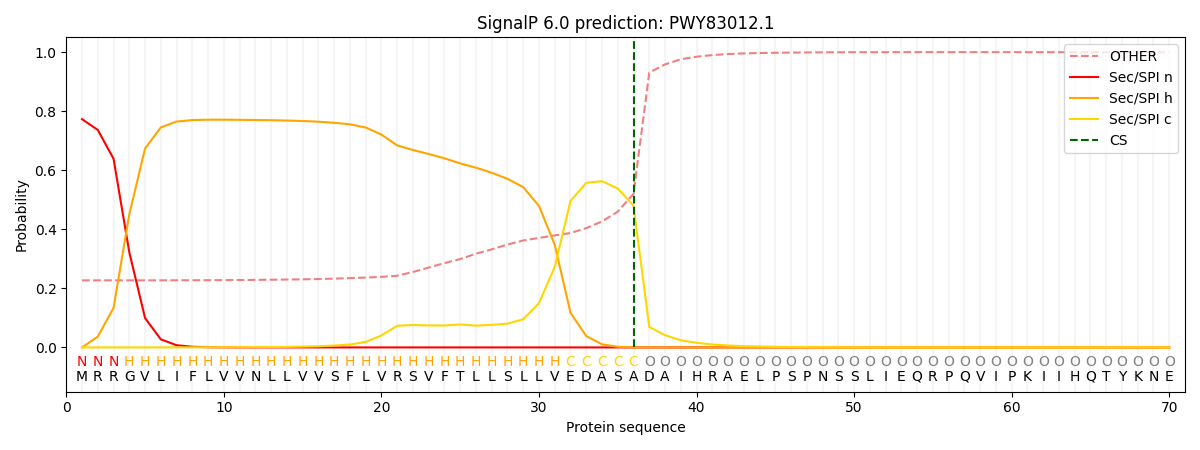
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.236192 | 0.763776 | CS pos: 36-37. Pr: 0.4799 |

