You are browsing environment: FUNGIDB
CAZyme Information: PWY82086.1
You are here: Home > Sequence: PWY82086.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus heteromorphus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus heteromorphus | |||||||||||
| CAZyme ID | PWY82086.1 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | beta-mannanase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.78:33 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 69 | 343 | 4.1e-86 | 0.9930795847750865 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 226444 | COG3934 | 3.71e-31 | 45 | 373 | 4 | 305 | Endo-1,4-beta-mannosidase [Carbohydrate transport and metabolism]. |
| 395098 | Cellulase | 6.24e-12 | 89 | 317 | 36 | 241 | Cellulase (glycosyl hydrolase family 5). |
| 226217 | XynA | 0.005 | 177 | 220 | 126 | 175 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.44e-228 | 1 | 378 | 1 | 381 | |
| 1.12e-225 | 36 | 378 | 18 | 360 | |
| 1.92e-225 | 1 | 377 | 1 | 383 | |
| 1.92e-225 | 1 | 377 | 1 | 383 | |
| 7.22e-222 | 36 | 378 | 18 | 360 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.27e-221 | 36 | 377 | 1 | 342 | The ligand-free structure of ManBK from Aspergillus niger BK01 [Aspergillus niger],3WH9_B The ligand-free structure of ManBK from Aspergillus niger BK01 [Aspergillus niger] |
|
| 1.01e-187 | 36 | 377 | 2 | 342 | Crtstal structure of glycoside hydrolase family 5 beta-mannanase from Talaromyces trachyspermus [Talaromyces trachyspermus] |
|
| 3.03e-154 | 36 | 378 | 3 | 344 | Chain A, ENDO-1,4-B-D-MANNANASE [Trichoderma reesei],1QNP_A Chain A, Endo-1,4-b-d-mannanase [Trichoderma reesei],1QNQ_A Chain A, Endo-1,4-b-d-mannanase [Trichoderma reesei],1QNR_A Chain A, Endo-1,4-b-d-mannanase [Trichoderma reesei],1QNS_A Chain A, Endo-1,4-b-d-mannanase [Trichoderma reesei] |
|
| 8.23e-149 | 41 | 378 | 19 | 355 | Chain A, Gh5 Endo-beta-1,4-mannanase [Podospora anserina] |
|
| 7.45e-81 | 36 | 380 | 3 | 359 | Crystal Structure Analysis of the Endo-1,4-beta-mannanase from Rhizomucor miehei [Rhizomucor miehei] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.27e-221 | 1 | 377 | 1 | 380 | Probable mannan endo-1,4-beta-mannosidase A OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=manA PE=1 SV=1 |
|
| 2.38e-210 | 1 | 381 | 1 | 376 | Mannan endo-1,4-beta-mannosidase A OS=Aspergillus aculeatus OX=5053 GN=manA PE=1 SV=1 |
|
| 1.52e-197 | 7 | 380 | 9 | 384 | Probable mannan endo-1,4-beta-mannosidase A-1 OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=manA-1 PE=3 SV=1 |
|
| 3.40e-195 | 37 | 380 | 26 | 369 | Probable mannan endo-1,4-beta-mannosidase A OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=manA PE=3 SV=1 |
|
| 6.96e-191 | 1 | 381 | 1 | 386 | Probable mannan endo-1,4-beta-mannosidase A OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=manA PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
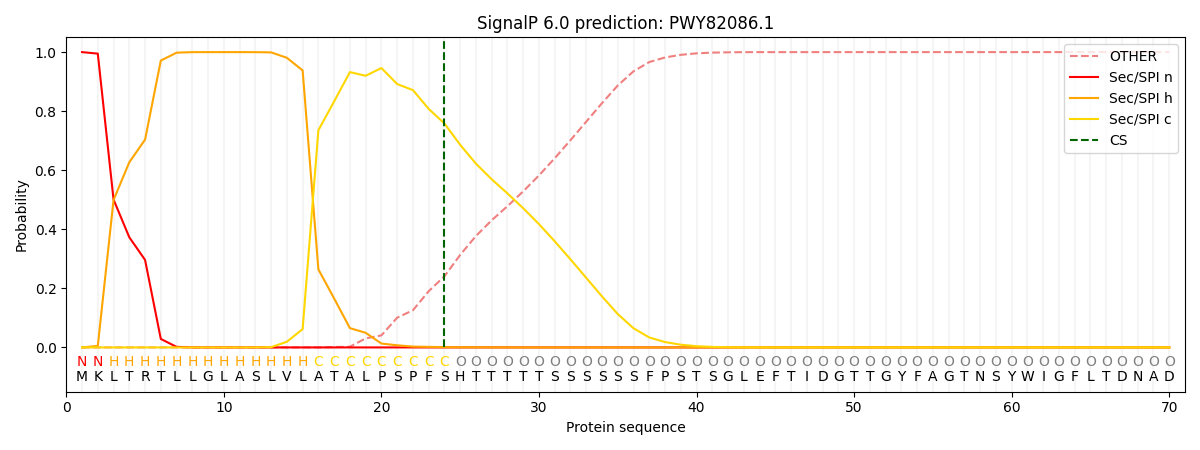
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000303 | 0.999657 | CS pos: 24-25. Pr: 0.7582 |
