You are browsing environment: FUNGIDB
CAZyme Information: PWY80559.1
You are here: Home > Sequence: PWY80559.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus heteromorphus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus heteromorphus | |||||||||||
| CAZyme ID | PWY80559.1 | |||||||||||
| CAZy Family | GH28 | |||||||||||
| CAZyme Description | glycoside hydrolase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.4:38 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 27 | 304 | 2.6e-111 | 0.9964539007092199 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395098 | Cellulase | 1.68e-72 | 31 | 297 | 2 | 272 | Cellulase (glycosyl hydrolase family 5). |
| 225344 | BglC | 2.31e-19 | 22 | 325 | 42 | 390 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
| 197593 | fCBD | 2.64e-13 | 374 | 405 | 2 | 33 | Fungal-type cellulose-binding domain. Small four-cysteine cellulose-binding domain of fungi |
| 395595 | CBM_1 | 3.27e-12 | 374 | 402 | 1 | 29 | Fungal cellulose binding domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.68e-217 | 3 | 405 | 4 | 409 | |
| 5.43e-216 | 3 | 405 | 4 | 409 | |
| 1.32e-214 | 3 | 405 | 4 | 410 | |
| 1.08e-213 | 3 | 405 | 4 | 410 | |
| 1.55e-192 | 1 | 405 | 1 | 411 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.07e-155 | 20 | 325 | 8 | 313 | Crystal structure of an endoglucanase from Penicillium verruculosum in complex with cellobiose [Talaromyces verruculosus],5L9C_B Crystal structure of an endoglucanase from Penicillium verruculosum in complex with cellobiose [Talaromyces verruculosus],5L9C_C Crystal structure of an endoglucanase from Penicillium verruculosum in complex with cellobiose [Talaromyces verruculosus],5L9C_D Crystal structure of an endoglucanase from Penicillium verruculosum in complex with cellobiose [Talaromyces verruculosus] |
|
| 8.61e-155 | 20 | 325 | 8 | 313 | CRYSTAL STRUCTURE OF EN ENDOGLUCANASE S308P FROM PENICILLIUM VERRUCULOSUM [Talaromyces verruculosus],6T9F_B CRYSTAL STRUCTURE OF EN ENDOGLUCANASE S308P FROM PENICILLIUM VERRUCULOSUM [Talaromyces verruculosus],6T9F_C CRYSTAL STRUCTURE OF EN ENDOGLUCANASE S308P FROM PENICILLIUM VERRUCULOSUM [Talaromyces verruculosus],6T9F_D CRYSTAL STRUCTURE OF EN ENDOGLUCANASE S308P FROM PENICILLIUM VERRUCULOSUM [Talaromyces verruculosus] |
|
| 1.72e-154 | 25 | 325 | 3 | 303 | Structure of Thermoascus aurantiacus family 5 endoglucanase [Thermoascus aurantiacus],1GZJ_B Structure of Thermoascus aurantiacus family 5 endoglucanase [Thermoascus aurantiacus] |
|
| 4.95e-154 | 20 | 325 | 8 | 313 | Chain A, Endoglucanase [Talaromyces verruculosus],6YON_B Chain B, Endoglucanase [Talaromyces verruculosus],6YON_C Chain C, Endoglucanase [Talaromyces verruculosus],6YON_D Chain D, Endoglucanase [Talaromyces verruculosus] |
|
| 9.95e-154 | 20 | 325 | 8 | 313 | CRYSTAL STRUCTURE OF AN ENDOGLUCANASE D213A FROM PENICILLIUM VERRUCULOSUM [Talaromyces verruculosus],6T9G_B CRYSTAL STRUCTURE OF AN ENDOGLUCANASE D213A FROM PENICILLIUM VERRUCULOSUM [Talaromyces verruculosus],6T9G_C CRYSTAL STRUCTURE OF AN ENDOGLUCANASE D213A FROM PENICILLIUM VERRUCULOSUM [Talaromyces verruculosus],6T9G_D CRYSTAL STRUCTURE OF AN ENDOGLUCANASE D213A FROM PENICILLIUM VERRUCULOSUM [Talaromyces verruculosus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.21e-151 | 9 | 329 | 6 | 325 | Endo-beta-1,4-glucanase A OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=eglA PE=1 SV=1 |
|
| 1.88e-145 | 20 | 325 | 27 | 331 | Probable endo-beta-1,4-glucanase B OS=Aspergillus kawachii (strain NBRC 4308) OX=1033177 GN=eglB PE=3 SV=1 |
|
| 3.95e-142 | 20 | 325 | 27 | 330 | Probable endo-beta-1,4-glucanase B OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=eglB PE=3 SV=1 |
|
| 3.95e-142 | 20 | 325 | 27 | 330 | Endo-beta-1,4-glucanase B OS=Aspergillus niger OX=5061 GN=eglB PE=1 SV=1 |
|
| 1.49e-141 | 19 | 325 | 21 | 327 | Probable endo-beta-1,4-glucanase B OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=eglB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
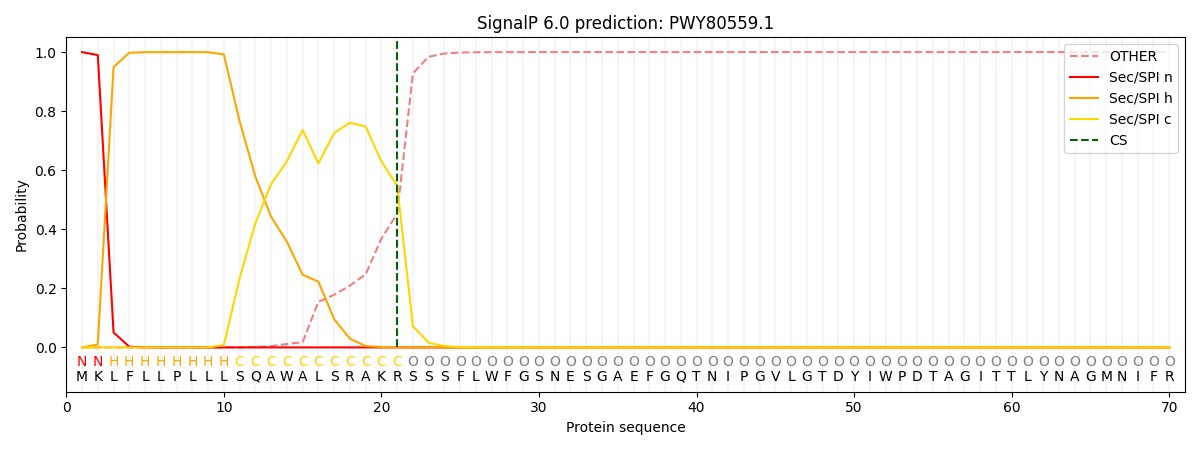
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000279 | 0.999697 | CS pos: 21-22. Pr: 0.5475 |
