You are browsing environment: FUNGIDB
CAZyme Information: PWY65731.1
You are here: Home > Sequence: PWY65731.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus heteromorphus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus heteromorphus | |||||||||||
| CAZyme ID | PWY65731.1 | |||||||||||
| CAZy Family | AA16 | |||||||||||
| CAZyme Description | pectin lyase 1 precursor | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 123016; End:124376 Strand: + | |||||||||||
Full Sequence Download help
| MKYTTILAAV ATFASTVSAG VTGSAEGFAK GVTGGGSATE VYPTTNEELV SYLGDSEARV | 60 |
| IVLDRTFDFT GTEGTTTTSG CSPWGTASSC QTAINLHNWC DNYEPNAPKV SSITYDTAGV | 120 |
| LGIIVNSNKS LVGKGSAGVI KGKGIRMVSG AKNIIIQNIA ITDINPQYVW GGDAITVDDA | 180 |
| DLVWIDHVTT ARIARQHIVL GTSADNRVTI SYSLIDGRSD YSATCNGHHY WGIYLDGSND | 240 |
| MVTLMGNYFY YLSGRMPKVQ GNTLLHAVNN LFHNIEGHAF EIGSGGYVLA EGNVFQNVDT | 300 |
| VVETPISGSL FSSPDATTNA QCKSVFGRSC QLNAFGSSGS WSGADTSIIS RFSGKNIASA | 360 |
| RAPTNIESWT MKNAGAGNL | 379 |
Enzyme Prediction help
| EC | 4.2.2.10:25 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL1 | 113 | 297 | 5.5e-86 | 0.9893048128342246 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 214765 | Amb_all | 2.26e-55 | 112 | 299 | 3 | 190 | Amb_all domain. |
| 366158 | Pec_lyase_C | 3.09e-17 | 123 | 295 | 30 | 211 | Pectate lyase. This enzyme forms a right handed beta helix structure. Pectate lyase is an enzyme involved in the maceration and soft rotting of plant tissue. |
| 226384 | PelB | 1.40e-15 | 8 | 379 | 20 | 344 | Pectate lyase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CAK37997.1|PL1_4|4.2.2.10 | 8.13e-222 | 8 | 377 | 9 | 378 |
| BCS05627.1|PL1_4 | 1.10e-219 | 8 | 377 | 9 | 378 |
| BCR92964.1|PL1_4 | 1.10e-219 | 8 | 377 | 9 | 378 |
| GAA92693.1|PL1_4|4.2.2.10 | 1.10e-219 | 8 | 377 | 9 | 378 |
| CAA01023.1|PL1_4|4.2.2.10 | 3.54e-218 | 8 | 377 | 9 | 377 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1QCX_A | 2.46e-219 | 19 | 377 | 1 | 358 | Pectin Lyase B [Aspergillus niger] |
| 1IDK_A | 3.04e-178 | 20 | 379 | 2 | 359 | Pectin Lyase A [Aspergillus niger] |
| 1IDJ_A | 6.12e-178 | 20 | 379 | 2 | 359 | Pectin Lyase A [Aspergillus niger],1IDJ_B Pectin Lyase A [Aspergillus niger] |
| 1AIR_A | 2.30e-11 | 148 | 313 | 107 | 275 | Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O88_A Chain A, Pectate Lyase C [Dickeya chrysanthemi],1O8D_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8E_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8F_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8G_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8H_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8I_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8J_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8K_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8L_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8M_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1PLU_A Chain A, Protein (pectate Lyase C) [Dickeya chrysanthemi],2PEC_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi] |
| 2EWE_A | 5.51e-11 | 148 | 313 | 107 | 275 | Chain A, Pectate lyase C [Dickeya chrysanthemi] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| sp|A2QFN7|PELB_ASPNC | 1.44e-222 | 8 | 377 | 9 | 378 | Probable pectin lyase B OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=pelB PE=3 SV=1 |
| sp|Q00205|PELB_ASPNG | 6.28e-219 | 8 | 377 | 9 | 377 | Pectin lyase B OS=Aspergillus niger OX=5061 GN=pelB PE=1 SV=1 |
| sp|Q4WV10|PELA_ASPFU | 1.60e-196 | 1 | 379 | 1 | 380 | Probable pectin lyase A OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=pelA PE=3 SV=1 |
| sp|B0XT36|PELA_ASPFC | 2.85e-194 | 1 | 379 | 1 | 378 | Probable pectin lyase A OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=pelA PE=3 SV=1 |
| sp|Q4WIS6|PELB_ASPFU | 2.85e-194 | 1 | 379 | 1 | 378 | Probable pectin lyase B OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=pelB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
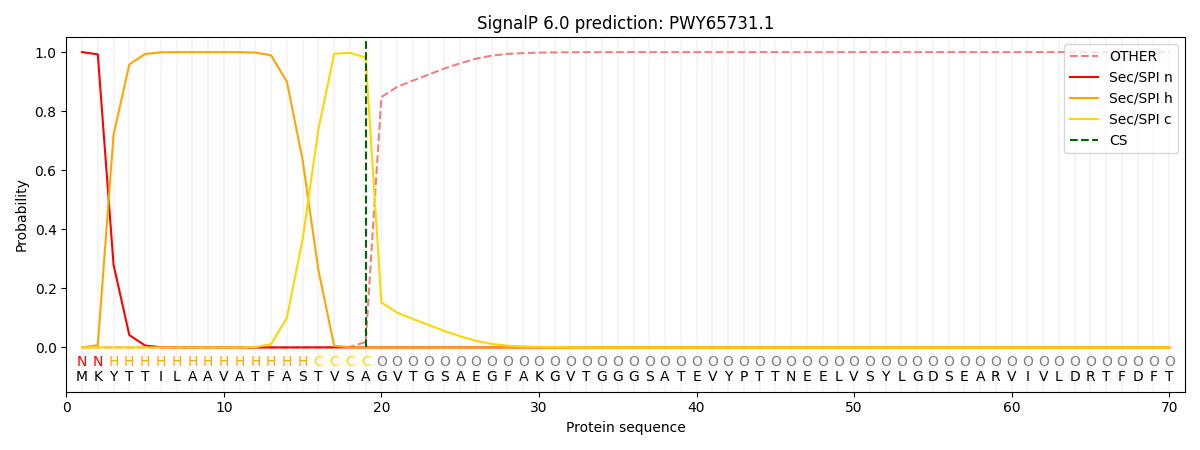
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000218 | 0.999739 | CS pos: 19-20. Pr: 0.9815 |
