You are browsing environment: FUNGIDB
CAZyme Information: PWY61812.1
You are here: Home > Sequence: PWY61812.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
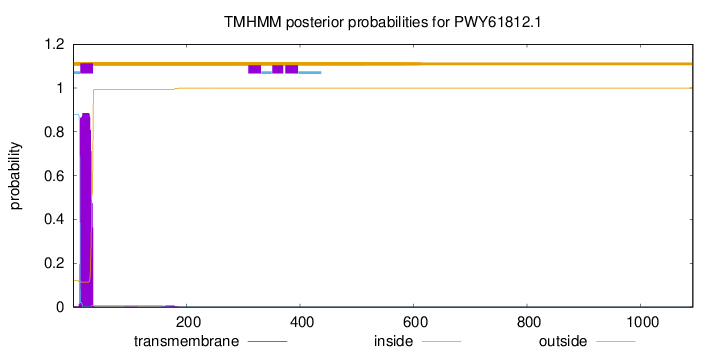
TMHMM annotations
Basic Information help
| Species | Aspergillus eucalypticola | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus eucalypticola | |||||||||||
| CAZyme ID | PWY61812.1 | |||||||||||
| CAZy Family | AA1 | |||||||||||
| CAZyme Description | ER glycosyl hydrolase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH47 | 50 | 602 | 1.4e-144 | 0.9955156950672646 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396217 | Glyco_hydro_47 | 2.04e-133 | 50 | 603 | 1 | 453 | Glycosyl hydrolase family 47. Members of this family are alpha-mannosidases that catalyze the hydrolysis of the terminal 1,2-linked alpha-D-mannose residues in the oligo-mannose oligosaccharide Man(9)(GlcNAc)(2). |
| 240427 | PTZ00470 | 1.71e-58 | 36 | 605 | 65 | 520 | glycoside hydrolase family 47 protein; Provisional |
| 238300 | PA | 0.002 | 965 | 1023 | 48 | 102 | PA: Protease-associated (PA) domain. The PA domain is an insert domain in a diverse fraction of proteases. The significance of the PA domain to many of the proteins in which it is inserted is undetermined. It may be a protein-protein interaction domain. At peptidase active sites, the PA domain may participate in substrate binding and/or promoting conformational changes, which influence the stability and accessibility of the site to substrate. Proteins into which the PA domain is inserted include the following: i) various signal peptide peptidases including, hSPPL2a and 2b which catalyze the intramembrane proteolysis of tumor necrosis factor alpha, ii) various proteins containing a C3H2C3 RING finger including, Arabidopsis ReMembR-H2 protein and various E3 ubiquitin ligases such as human GRAIL (gene related to anergy in lymphocytes), iii) EDEM3 (ER-degradation-enhancing mannosidase-like 3 protein), iv) various plant vacuolar sorting receptors such as Pisum sativum BP-80, v) glutamate carboxypeptidase II (GCPII), vi) yeast aminopeptidase Y, vii) Vibrio metschnikovii VapT, a sodium dodecyl sulfate (SDS) resistant extracellular alkaline serine protease, viii) lactocepin (a cell envelope-associated protease from Lactobacillus paracasei subsp. paracasei NCDO 151), ix) various subtilisin-like proteases such as melon Cucumisin, and x) human TfR (transferrin receptor) 1 and 2. |
| 239038 | PA_C_RZF_like | 0.002 | 910 | 1016 | 3 | 115 | PA_C-RZF_ like: Protease-associated (PA) domain C_RZF-like. This group includes various PA domain-containing proteins similar to C-RZF (chicken embryo RING zinc finger) protein. These proteins contain a C3H2C3 RING finger. C-RZF is expressed in embryo cells and is restricted mainly to brain and heart, it is localized to both the nucleus and endosomes. Additional C3H2C3 RING finger proteins belonging to this group, include Arabidopsis ReMembR-H2 protein and mouse sperizin. ReMembR-H2 is likely to be an integral membrane protein, and to traffic through the endosomal pathway. Sperizin is expressed in haploid germ cells and localized in the cytoplasm, it may participate in spermatogenesis. The significance of the PA domain to these proteins has not been ascertained. It may be a protein-protein interaction domain. At peptidase active sites, the PA domain may participate in substrate binding and/or promoting conformational changes, which influence the stability and accessibility of the site to substrate. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 1092 | 1 | 1096 | |
| 0.0 | 1 | 1092 | 1 | 1096 | |
| 0.0 | 44 | 1092 | 34 | 1083 | |
| 0.0 | 19 | 1092 | 1 | 965 | |
| 0.0 | 1 | 1092 | 1 | 1090 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.49e-30 | 50 | 603 | 12 | 451 | Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase and Man9GlcNAc2-PA complex [Homo sapiens] |
|
| 1.62e-30 | 50 | 603 | 17 | 456 | Crystal Structure Of Human Class I Alpha1,2-Mannosidase [Homo sapiens] |
|
| 1.67e-30 | 50 | 603 | 17 | 456 | Crystal Structure Of Human Class I Alpha1,2-Mannosidase In Complex With 1-Deoxymannojirimycin [Homo sapiens],1FO3_A Crystal Structure Of Human Class I Alpha1,2-Mannosidase In Complex With Kifunensine [Homo sapiens] |
|
| 4.33e-30 | 50 | 603 | 95 | 534 | Crystal Structure Of Human Class I alpha-1,2-Mannosidase In Complex With Thio-Disaccharide Substrate Analogue [Homo sapiens] |
|
| 6.80e-30 | 50 | 603 | 12 | 451 | Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase T688A mutant and Thio-disaccharide substrate analog complex [Homo sapiens],5KK7_B Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase T688A mutant and Thio-disaccharide substrate analog complex [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.32e-119 | 34 | 614 | 26 | 488 | ER degradation-enhancing alpha-mannosidase-like protein 1 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=mnl1 PE=3 SV=2 |
|
| 1.63e-110 | 42 | 605 | 124 | 583 | ER degradation-enhancing alpha-mannosidase-like protein 1 OS=Mus musculus OX=10090 GN=Edem1 PE=1 SV=1 |
|
| 3.64e-110 | 42 | 599 | 129 | 582 | ER degradation-enhancing alpha-mannosidase-like protein 1 OS=Homo sapiens OX=9606 GN=EDEM1 PE=1 SV=1 |
|
| 3.26e-102 | 32 | 676 | 24 | 572 | ER degradation-enhancing alpha-mannosidase-like protein 1 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=MNL1 PE=1 SV=1 |
|
| 1.91e-90 | 31 | 610 | 28 | 481 | Alpha-mannosidase I MNS4 OS=Arabidopsis thaliana OX=3702 GN=MNS4 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
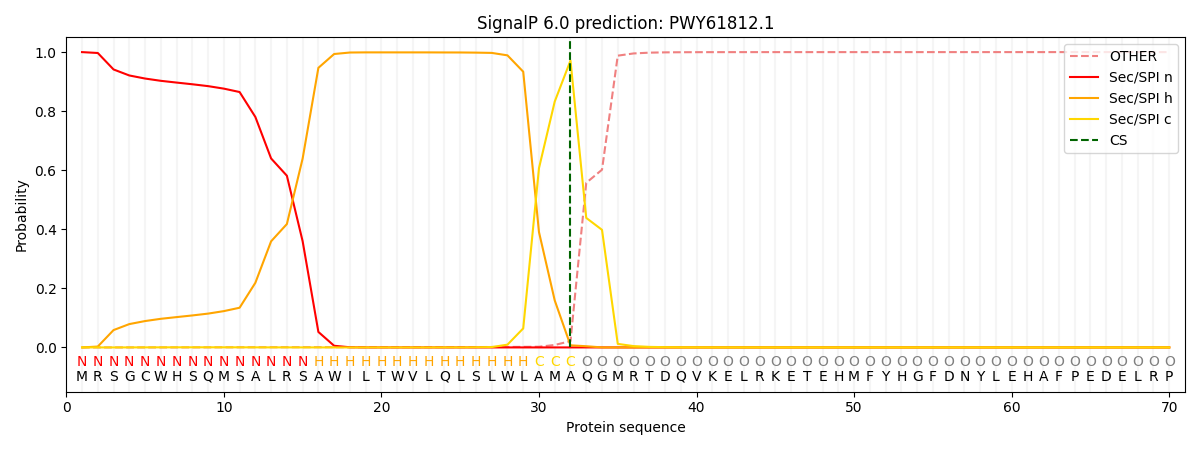
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000309 | 0.999657 | CS pos: 32-33. Pr: 0.9715 |