You are browsing environment: FUNGIDB
CAZyme Information: PV10_03052-t45_1-p1
You are here: Home > Sequence: PV10_03052-t45_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Exophiala mesophila | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Herpotrichiellaceae; Exophiala; Exophiala mesophila | |||||||||||
| CAZyme ID | PV10_03052-t45_1-p1 | |||||||||||
| CAZy Family | GH128 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH79 | 42 | 489 | 1.6e-57 | 0.9120879120879121 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 407104 | Glyco_hydro_79C | 7.35e-22 | 421 | 525 | 2 | 103 | Glycosyl hydrolase family 79 C-terminal beta domain. This domain is found at the C-terminus of glycosyl hydrolase family 79 proteins. It's function is not yet known. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.43e-181 | 11 | 545 | 11 | 548 | |
| 8.28e-177 | 17 | 533 | 18 | 531 | |
| 5.78e-172 | 1 | 533 | 1 | 530 | |
| 1.43e-171 | 16 | 533 | 18 | 533 | |
| 3.19e-168 | 14 | 533 | 312 | 832 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.68e-17 | 24 | 455 | 3 | 388 | Chain A, 4-O-alpha-L-rhamnosyl-beta-D-glucuronidase [Fusarium oxysporum],7DFS_A Chain A, 4-O-alpha-L-rhamnosyl-beta-D-glucuronidase [Fusarium oxysporum] |
|
| 1.23e-10 | 84 | 409 | 75 | 372 | beta-glucuronidase with an activity-based probe (N-acyl cyclophellitol aziridine) bound [Acidobacterium capsulatum],5G0Q_A beta-glucuronidase with an activity-based probe (N-alkyl cyclophellitol aziridine) bound [Acidobacterium capsulatum] |
|
| 1.27e-10 | 84 | 409 | 75 | 372 | Crystal structure of beta-glucuronidase from Acidobacterium capsulatum [Acidobacterium capsulatum ATCC 51196],3VNZ_A Crystal structure of beta-glucuronidase from Acidobacterium capsulatum in complex with D-glucuronic acid [Acidobacterium capsulatum ATCC 51196],3VO0_A Crystal structure of beta-glucuronidase from Acidobacterium capsulatum covalent-bonded with 2-deoxy-2-fluoro-D-glucuronic acid [Acidobacterium capsulatum ATCC 51196] |
|
| 2.97e-10 | 84 | 409 | 95 | 392 | A glycoside hydrolase mutant with an unreacted activity based probe bound [Acidobacterium capsulatum] |
|
| 2.90e-07 | 19 | 529 | 10 | 446 | Crystallographic structure of a bacterial heparanase [Burkholderia pseudomallei],5BWI_B Crystallographic structure of a bacterial heparanase [Burkholderia pseudomallei] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.12e-38 | 25 | 528 | 34 | 541 | Beta-glucuronidase OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=An02g11890 PE=1 SV=1 |
|
| 1.37e-34 | 16 | 506 | 12 | 525 | Beta-glucuronidase OS=Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) OX=367110 GN=NCU00937 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
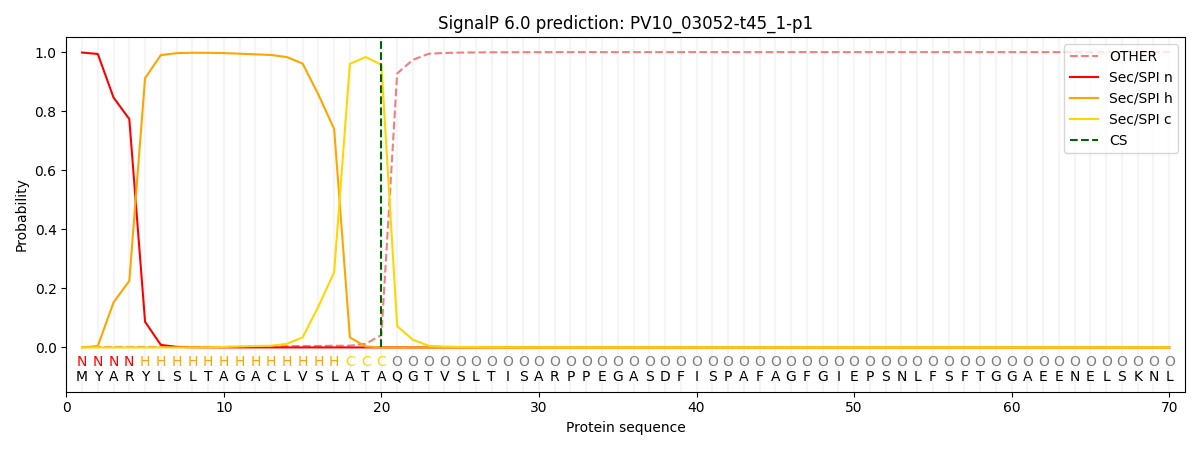
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.002994 | 0.996994 | CS pos: 20-21. Pr: 0.9573 |
