You are browsing environment: FUNGIDB
CAZyme Information: PV06_01020-t43_2-p1
You are here: Home > Sequence: PV06_01020-t43_2-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Exophiala oligosperma | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Herpotrichiellaceae; Exophiala; Exophiala oligosperma | |||||||||||
| CAZyme ID | PV06_01020-t43_2-p1 | |||||||||||
| CAZy Family | AA1 | |||||||||||
| CAZyme Description | hypothetical protein, variant | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH12 | 85 | 213 | 1.3e-25 | 0.8141025641025641 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396303 | Glyco_hydro_12 | 1.73e-15 | 16 | 221 | 29 | 206 | Glycosyl hydrolase family 12. |
| 235746 | PRK06215 | 0.004 | 110 | 212 | 132 | 228 | hypothetical protein; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 5.74e-67 | 1 | 223 | 99 | 324 | |
| 5.74e-67 | 1 | 223 | 99 | 324 | |
| 3.25e-66 | 1 | 223 | 99 | 324 | |
| 1.34e-65 | 1 | 221 | 99 | 320 | |
| 1.34e-65 | 1 | 221 | 99 | 320 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.16e-18 | 16 | 221 | 47 | 226 | The X-ray Crystal Structure of the Trichoderma harzianum Endoglucanase 3 from family GH12 [Trichoderma harzianum],4H7M_B The X-ray Crystal Structure of the Trichoderma harzianum Endoglucanase 3 from family GH12 [Trichoderma harzianum] |
|
| 1.34e-14 | 17 | 220 | 41 | 218 | Chain A, ENDO-BETA-1-4-GLUCANASE [Trichoderma citrinoviride],1OA3_B Chain B, ENDO-BETA-1-4-GLUCANASE [Trichoderma citrinoviride],1OA3_C Chain C, ENDO-BETA-1-4-GLUCANASE [Trichoderma citrinoviride],1OA3_D Chain D, ENDO-BETA-1-4-GLUCANASE [Trichoderma citrinoviride] |
|
| 1.73e-12 | 20 | 220 | 44 | 218 | Chain A, ENDO-BETA-1,4-GLUCANASE [Trichoderma reesei],1OLQ_B Chain B, ENDO-BETA-1,4-GLUCANASE [Trichoderma reesei] |
|
| 2.39e-12 | 20 | 220 | 44 | 218 | Chain A, ENDO-BETA-1,4-GLUCANASE [Trichoderma reesei],1H8V_B Chain B, ENDO-BETA-1,4-GLUCANASE [Trichoderma reesei],1H8V_C Chain C, ENDO-BETA-1,4-GLUCANASE [Trichoderma reesei],1H8V_D Chain D, ENDO-BETA-1,4-GLUCANASE [Trichoderma reesei],1H8V_E Chain E, ENDO-BETA-1,4-GLUCANASE [Trichoderma reesei],1H8V_F Chain F, ENDO-BETA-1,4-GLUCANASE [Trichoderma reesei] |
|
| 2.39e-12 | 20 | 220 | 44 | 218 | Chain A, ENDO-BETA-1,4-GLUCANASE [Trichoderma reesei],1OA2_B Chain B, ENDO-BETA-1,4-GLUCANASE [Trichoderma reesei],1OA2_C Chain C, ENDO-BETA-1,4-GLUCANASE [Trichoderma reesei],1OA2_D Chain D, ENDO-BETA-1,4-GLUCANASE [Trichoderma reesei],1OA2_E Chain E, ENDO-BETA-1,4-GLUCANASE [Trichoderma reesei],1OA2_F Chain F, ENDO-BETA-1,4-GLUCANASE [Trichoderma reesei] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.24e-17 | 20 | 207 | 66 | 225 | Xyloglucan-specific endo-beta-1,4-glucanase A OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=xgeA PE=1 SV=1 |
|
| 1.94e-15 | 20 | 221 | 67 | 239 | Probable xyloglucan-specific endo-beta-1,4-glucanase A OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=xgeA PE=3 SV=1 |
|
| 1.94e-15 | 20 | 221 | 67 | 239 | Probable xyloglucan-specific endo-beta-1,4-glucanase A OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=xgeA PE=3 SV=1 |
|
| 4.59e-12 | 20 | 207 | 67 | 227 | Xyloglucan-specific endo-beta-1,4-glucanase A OS=Aspergillus niger OX=5061 GN=xgeA PE=1 SV=1 |
|
| 4.59e-12 | 20 | 207 | 67 | 227 | Probable xyloglucan-specific endo-beta-1,4-glucanase A OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=xgeA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
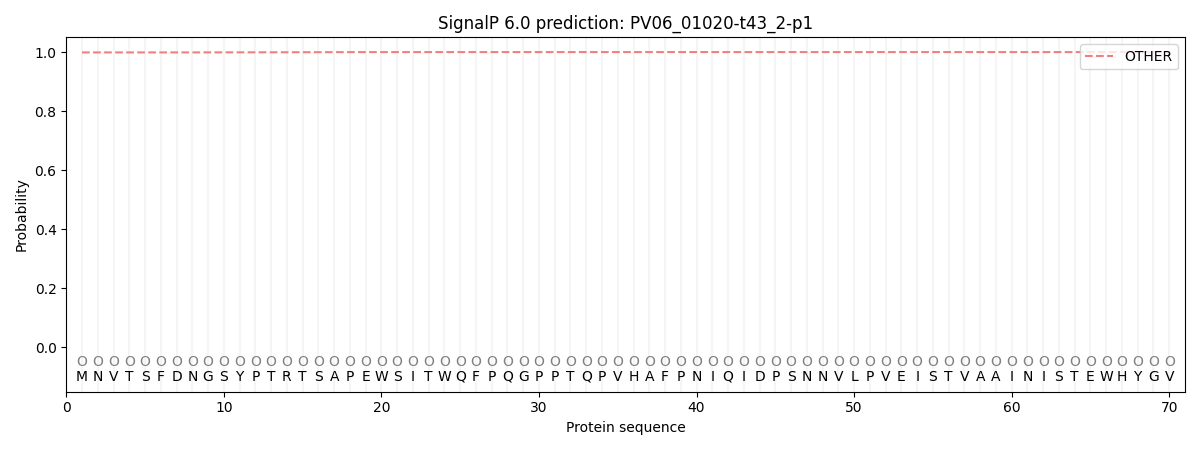
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.999022 | 0.001035 |
