You are browsing environment: FUNGIDB
CAZyme Information: PUG3_T011719-RA-p1
You are here: Home > Sequence: PUG3_T011719-RA-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Globisporangium ultimum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Pythiaceae; Globisporangium; Globisporangium ultimum | |||||||||||
| CAZyme ID | PUG3_T011719-RA-p1 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | Sterol 3-beta-glucosyltransferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 2.4.1.173:23 | 2.4.1.-:3 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT1 | 14 | 321 | 2.2e-18 | 0.6910994764397905 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 340817 | GT1_Gtf-like | 4.13e-14 | 1 | 322 | 110 | 401 | UDP-glycosyltransferases and similar proteins. This family includes the Gtfs, a group of homologous glycosyltransferases involved in the final stages of the biosynthesis of antibiotics vancomycin and related chloroeremomycin. Gtfs transfer sugar moieties from an activated NDP-sugar donor to the oxidatively cross-linked heptapeptide core of vancomycin group antibiotics. The core structure is important for the bioactivity of the antibiotics. |
| 224732 | YjiC | 1.33e-10 | 219 | 320 | 292 | 393 | UDP:flavonoid glycosyltransferase YjiC, YdhE family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.01e-81 | 2 | 469 | 946 | 1436 | |
| 5.72e-57 | 3 | 481 | 928 | 1402 | |
| 5.55e-42 | 3 | 342 | 215 | 535 | |
| 2.55e-41 | 3 | 342 | 215 | 535 | |
| 9.80e-41 | 3 | 351 | 355 | 677 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.01e-13 | 6 | 326 | 134 | 439 | Sterol 3-beta-glucosyltransferase (ugt51) from Saccharomyces cerevisiae (strain ATCC 204508 / S288c) [Saccharomyces cerevisiae S288C],5XVM_B Sterol 3-beta-glucosyltransferase (ugt51) from Saccharomyces cerevisiae (strain ATCC 204508 / S288c) [Saccharomyces cerevisiae S288C] |
|
| 3.17e-13 | 6 | 326 | 154 | 459 | Sterol 3-beta-glucosyltransferase (ugt51) from Saccharomyces cerevisiae (strain ATCC 204508 / S288c): UDPG complex [Saccharomyces cerevisiae S288C],5GL5_B Sterol 3-beta-glucosyltransferase (ugt51) from Saccharomyces cerevisiae (strain ATCC 204508 / S288c): UDPG complex [Saccharomyces cerevisiae S288C] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.42e-32 | 3 | 327 | 274 | 575 | Sterol 3-beta-glucosyltransferase UGT80B1 OS=Arabidopsis thaliana OX=3702 GN=UGT80B1 PE=1 SV=1 |
|
| 1.32e-27 | 3 | 327 | 311 | 611 | Sterol 3-beta-glucosyltransferase UGT80A2 OS=Arabidopsis thaliana OX=3702 GN=UGT80A2 PE=1 SV=1 |
|
| 3.02e-18 | 6 | 326 | 848 | 1153 | Sterol 3-beta-glucosyltransferase OS=Kluyveromyces lactis (strain ATCC 8585 / CBS 2359 / DSM 70799 / NBRC 1267 / NRRL Y-1140 / WM37) OX=284590 GN=ATG26 PE=3 SV=1 |
|
| 1.64e-15 | 3 | 327 | 1276 | 1610 | UDP-sugar-dependent glycosyltransferase 52 OS=Dictyostelium discoideum OX=44689 GN=ugt52 PE=2 SV=1 |
|
| 1.98e-14 | 6 | 326 | 1127 | 1433 | Sterol 3-beta-glucosyltransferase OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=ATG26 PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
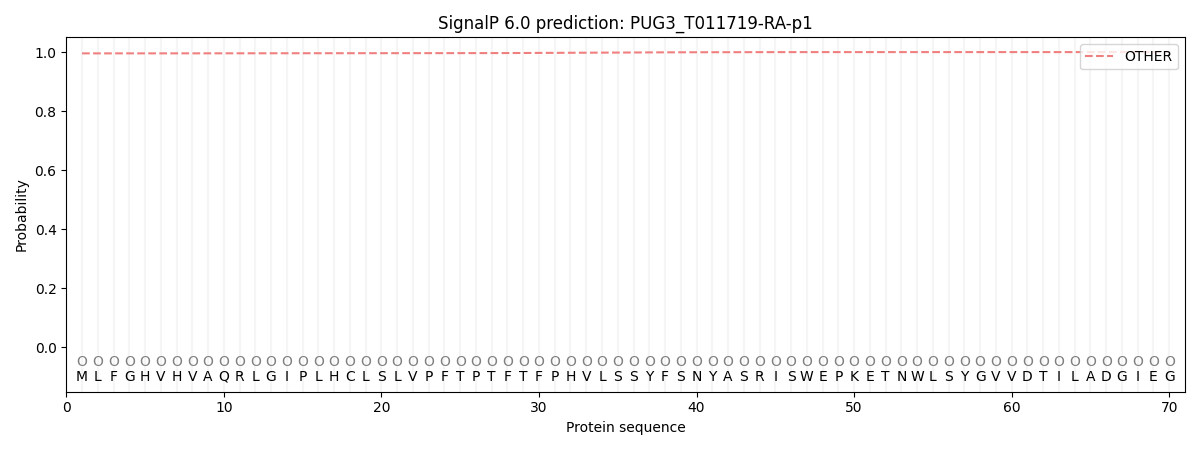
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.996117 | 0.003920 |
