You are browsing environment: FUNGIDB
CAZyme Information: PTTG_27250-t43_1-p1
You are here: Home > Sequence: PTTG_27250-t43_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Puccinia triticina | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Basidiomycota; Pucciniomycetes; ; Pucciniaceae; Puccinia; Puccinia triticina | |||||||||||
| CAZyme ID | PTTG_27250-t43_1-p1 | |||||||||||
| CAZy Family | GT39 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 338242; End:346803 Strand: - | |||||||||||
Full Sequence Download help
| MATITISSRT RLICLTILLL GLQLSLIHSY PYDPAFSDYN ININSSAGNN ILDYYSDWPE | 60 |
| KSQNGSRYTP SPANWRELPI YTVILDRFMD GNPSNNDFYK SRSEWDFLSN QLRHGGDIEG | 120 |
| FAQDRVLDYI YGMGYRTIYI AGTPWLNMPW QSDGYSALDF TLLDPHFGTL SEWRAAIDKI | 180 |
| HSKGMYVMLD TTTTTLSDFL EFKGNSGGAA PFNLHGYEVE YKTTVQKPWN INQYAEFKFT | 240 |
| NSRDQSCRLP KFYNPNGTDV LPPTDWDGCY AGDFNQFGDS SPAGKAPGWQ DQLTKYSGVQ | 300 |
| DRLRDWDSGV AAKLEKLACM TVKALDMDAL RVDKATQQTL EFMGKWGGSL RTCAKDLGKN | 360 |
| NFFITGEISD GNTFGSLYIG RGRQPAHYAN LEFDTAALVT ANQTENFMRP IGENALDSAA | 420 |
| FHYSIYRSLT RFLGMDGELD SPYDIPVNWV EAWRQIFVTN DLFNQETQQL DPRHLYGTTN | 480 |
| QDNFRWASLI NGTERFLLGQ LVTNLIMPGI PFAFYGEEQD LHIFDSQAND YLYGRQPMSS | 540 |
| SKAWQTHGCY KLGSKKYPTM PLGKALNGCK DDWNSLDHYD PTSNTRNMLA HFAYLRSQYA | 600 |
| SLQDGFNLTQ LGNWTHFGEL PASSHTPTEW GLWSVSRSSL KSQNLSGPNS QLPVWILYSN | 660 |
| VNQTTTFSYD CSSKLAITSP YPGPSTVRNL FYPYETYDLD PSSTSSTWDD APPFLGCLKS | 720 |
| ITMDPLSYKV LVPASNWVSP QPRLVGFVPG HDARILSQAD NDNETIPISL SFSDELSCEG | 780 |
| VSSSLSLSYT IDPNSKASPR IDLEHANCSK IESPSNSTQS ALPAVWQWSA SIVGAADGIY | 840 |
| ELILNNATGQ NGIHTNSIDH LLLRKGRKLN PITFSDVSYS ESLLENKDGK YTLRSDAPGA | 900 |
| DMMRYSVDFG KTYSNWTRYS SDHVRVQYWS RLTGSAAPTI DADINFDGGF KRRFPQLILR | 960 |
| GAYNKWGFDE GIPGLLSPRS ENMTIDVITS WPHEFQLAVF EARDKVFYGD VDNDGVLDLL | 1020 |
| PPNSQARNFL SLPPPSKPYL GWRIMINPRD LTWGAQPVGH QKIVIMLLFL LLFIPPTTAL | 1080 |
| LACWLYQRIF YKIKHNQYGF QTAGLNSFLP FQHLPKSMLN SQIPSTLKHA ITPQLGSKKE | 1140 |
| TPHFSFANTW PENINSRRKV LIATLEYEII DWGIKVKIGG LGVMTSLMGK VMEDLDLLWV | 1200 |
| IPKVSGLEYP EAERAEPIPV VVFGEKYLVE LQIHKFKNIT YFLLDSPIFR ANSKMNPYPA | 1260 |
| RMDDLDSAIF YSYWNQSIAE ICRRTPDLTI YHINDYHGAL APLYLLPKII PVALSLHNAE | 1320 |
| FQGQWPLGTP EEENEVCQSF NISSKVCSKY ARYGSVFNLL HSAASYISHH QNSVGVAGVS | 1380 |
| EKYGKRSWAR YPVLWTLKSI EPLPNPDPTD VESLDVIPPD MRRIQIDQEA EAKRPGNKLK | 1440 |
| LQEWAGLHED PKAQVFVFVG RWSFQKGVDL IADVFPLLLE TRKDVQLVAI GPVIDLYGKL | 1500 |
| AALKLNRLME SYPDRVYSKP EFTALPDFRD EPFGLVAVEF GRKGALGVGS RLGGLGLMPG | 1560 |
| WWFPVESDST AHLHSQFAKT LNAALECPEE ERAVLRARSV HQRFPVLEWR MKMEDIHSRS | 1620 |
| VKASRKYAGR LTMNINQNEV IKPDESSSAS HDQGNPRQAH LSATESNKPG QICTSSTGNQ | 1680 |
| SNTFGSHLNP TRPSSDVSRI LSSTNQLAVQ LSEAVKRLII TKNRSSTPPS RLSGRITDKA | 1740 |
| SVDGAITPSG RPQNNDQEIL TSVVTLDKNN GNTGEHNPAD FQRTYCGQDD RNSPLNQGVK | 1800 |
| DFTDEDGKAS QEFVRRLQTL DASNSQRELC IARYIVAAHK AHFNQVRKGT LALAKTKYGS | 1860 |
| SPTLSVTSII PRAMLGISSH ANDLHDFSDQ VKNEKNDTVD VTAALSMNRW QIRLQRRVFG | 1920 |
| WPVYTVLLAI GQILGATSFQ LSLLSGTSSN GSFDFYVIGA VNILGSAWWY GWSRKKPSTW | 1980 |
| SLSMPWIFFG LAFIFIGLPS LSENLKQYSR RHSLAVAASG FYSFASSAGF LFFSSNFGRR | 2040 |
| PLVLGIQTPK RDPTNAALAP PGWINAITLS LGAGCFSIAY ILFTGLPKYY RNVPGTVPNF | 2100 |
| TRALFRRKLV LWYLAAEILR NYWLSGPYGR NWNFLWTRET SFGVTLFLLL FFFVGVWGLA | 2160 |
| IWGLSRASKS HTWIVAIFAI GLGAPRWCQM LWGTSSVASY ISWGGSAGSH IATSLWLWLG | 2220 |
| VLDAVQGVGL GMILLQTLSR VHVAATLCLA QIIGSTAVLV ARATAPDRIG PGGVFPDLAL | 2280 |
| WNPKFSYLDS PLANWQFWIA LICQLIIVTG YFVLFRREQL SKP | 2323 |
Enzyme Prediction help
| EC | 2.4.1.183:18 | 2.4.1.-:2 | 2.4.1.183:36 | 2.4.1.-:11 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH13 | 115 | 519 | 7.5e-145 | 0.995 |
| GH13 | 1159 | 1526 | 8.9e-62 | 0.7775 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 200462 | AmyAc_AGS | 0.0 | 31 | 599 | 16 | 569 | Alpha amylase catalytic domain found in Alpha 1,3-glucan synthase (also called uridine diphosphoglucose-1,3-alpha-glucan glucosyltransferase and 1,3-alpha-D-glucan synthase). Alpha 1,3-glucan synthase (AGS, EC 2.4.1.183) is an enzyme that catalyzes the reversible chemical reaction of UDP-glucose and [alpha-D-glucosyl-(1-3)]n to form UDP and [alpha-D-glucosyl-(1-3)]n+1. AGS is a component of fungal cell walls. The cell wall of filamentous fungi is composed of 10-15% chitin and 10-35% alpha-1,3-glucan. AGS is triggered in fungi as a response to cell wall stress and elongates the glucan chains in cell wall synthesis. This group includes proteins from Ascomycetes and Basidomycetes. The Alpha-amylase family comprises the largest family of glycoside hydrolases (GH), with the majority of enzymes acting on starch, glycogen, and related oligo- and polysaccharides. These proteins catalyze the transformation of alpha-1,4 and alpha-1,6 glucosidic linkages with retention of the anomeric center. The protein is described as having 3 domains: A, B, C. A is a (beta/alpha) 8-barrel; B is a loop between the beta 3 strand and alpha 3 helix of A; C is the C-terminal extension characterized by a Greek key. The majority of the enzymes have an active site cleft found between domains A and B where a triad of catalytic residues (Asp, Glu and Asp) performs catalysis. Other members of this family have lost the catalytic activity as in the case of the human 4F2hc, or only have 2 residues that serve as the catalytic nucleophile and the acid/base, such as Thermus A4 beta-galactosidase with 2 Glu residues (GH42) and human alpha-galactosidase with 2 Asp residues (GH31). The family members are quite extensive and include: alpha amylase, maltosyltransferase, cyclodextrin glycotransferase, maltogenic amylase, neopullulanase, isoamylase, 1,4-alpha-D-glucan maltotetrahydrolase, 4-alpha-glucotransferase, oligo-1,6-glucosidase, amylosucrase, sucrose phosphorylase, and amylomaltase. |
| 340822 | GT5_Glycogen_synthase_DULL1-like | 7.34e-91 | 1159 | 1604 | 1 | 459 | Glycogen synthase GlgA and similar proteins. This family is most closely related to the GT5 family of glycosyltransferases. Glycogen synthase (EC:2.4.1.21) catalyzes the formation and elongation of the alpha-1,4-glucose backbone using ADP-glucose, the second and key step of glycogen biosynthesis. This family includes starch synthases of plants, such as DULL1 in Zea mays and glycogen synthases of various organisms. |
| 223443 | AmyA | 3.30e-25 | 80 | 618 | 3 | 464 | Glycosidase [Carbohydrate transport and metabolism]. |
| 200478 | AmyAc_bac_CMD_like_2 | 7.77e-21 | 76 | 190 | 1 | 118 | Alpha amylase catalytic domain found in bacterial cyclomaltodextrinases and related proteins. Cyclomaltodextrinase (CDase; EC3.2.1.54), neopullulanase (NPase; EC 3.2.1.135), and maltogenic amylase (MA; EC 3.2.1.133) catalyze the hydrolysis of alpha-(1,4) glycosidic linkages on a number of substrates including cyclomaltodextrins (CDs), pullulan, and starch. These enzymes hydrolyze CDs and starch to maltose and pullulan to panose by cleavage of alpha-1,4 glycosidic bonds whereas alpha-amylases essentially lack activity on CDs and pullulan. They also catalyze transglycosylation of oligosaccharides to the C3-, C4- or C6-hydroxyl groups of various acceptor sugar molecules. Since these proteins are nearly indistinguishable from each other, they are referred to as cyclomaltodextrinases (CMDs). This group of CMDs is bacterial. The Alpha-amylase family comprises the largest family of glycoside hydrolases (GH), with the majority of enzymes acting on starch, glycogen, and related oligo- and polysaccharides. These proteins catalyze the transformation of alpha-1,4 and alpha-1,6 glucosidic linkages with retention of the anomeric center. The protein is described as having 3 domains: A, B, C. A is a (beta/alpha) 8-barrel; B is a loop between the beta 3 strand and alpha 3 helix of A; C is the C-terminal extension characterized by a Greek key. The majority of the enzymes have an active site cleft found between domains A and B where a triad of catalytic residues (Asp, Glu and Asp) performs catalysis. Other members of this family have lost the catalytic activity as in the case of the human 4F2hc, or only have 2 residues that serve as the catalytic nucleophile and the acid/base, such as Thermus A4 beta-galactosidase with 2 Glu residues (GH42) and human alpha-galactosidase with 2 Asp residues (GH31). The family members are quite extensive and include: alpha amylase, maltosyltransferase, cyclodextrin glycotransferase, maltogenic amylase, neopullulanase, isoamylase, 1,4-alpha-D-glucan maltotetrahydrolase, 4-alpha-glucotransferase, oligo-1,6-glucosidase, amylosucrase, sucrose phosphorylase, and amylomaltase. |
| 200459 | AmyAc_AmyMalt_CGTase_like | 3.66e-19 | 80 | 540 | 7 | 369 | Alpha amylase catalytic domain found in maltogenic amylases, cyclodextrin glycosyltransferase, and related proteins. Enzymes such as amylases, cyclomaltodextrinase (CDase), and cyclodextrin glycosyltransferase (CGTase) degrade starch to smaller oligosaccharides by hydrolyzing the alpha-D-(1,4) linkages between glucose residues. In the case of CGTases, an additional cyclization reaction is catalyzed yielding mixtures of cyclic oligosaccharides which are referred to as alpha-, beta-, or gamma-cyclodextrins (CDs), consisting of six, seven, or eight glucose residues, respectively. CGTases are characterized depending on the major product of the cyclization reaction. Besides having similar catalytic site residues, amylases and CGTases contain carbohydrate binding domains that are distant from the active site and are implicated in attaching the enzyme to raw starch granules and in guiding the amylose chain into the active site. The maltogenic alpha-amylase from Bacillus is a five-domain structure, unlike most alpha-amylases, but similar to that of cyclodextrin glycosyltransferase. In addition to the A, B, and C domains, they have a domain D and a starch-binding domain E. Maltogenic amylase is an endo-acting amylase that has activity on cyclodextrins, terminally modified linear maltodextrins, and amylose. The Alpha-amylase family comprises the largest family of glycoside hydrolases (GH), with the majority of enzymes acting on starch, glycogen, and related oligo- and polysaccharides. These proteins catalyze the transformation of alpha-1,4 and alpha-1,6 glucosidic linkages with retention of the anomeric center. The protein is described as having 3 domains: A, B, C. A is a (beta/alpha) 8-barrel; B is a loop between the beta 3 strand and alpha 3 helix of A; C is the C-terminal extension characterized by a Greek key. The majority of the enzymes have an active site cleft found between domains A and B where a triad of catalytic residues (Asp, Glu and Asp) performs catalysis. Other members of this family have lost the catalytic activity as in the case of the human 4F2hc, or only have 2 residues that serve as the catalytic nucleophile and the acid/base, such as Thermus A4 beta-galactosidase with 2 Glu residues (GH42) and human alpha-galactosidase with 2 Asp residues (GH31). The family members are quite extensive and include: alpha amylase, maltosyltransferase, cyclodextrin glycotransferase, maltogenic amylase, neopullulanase, isoamylase, 1,4-alpha-D-glucan maltotetrahydrolase, 4-alpha-glucotransferase, oligo-1,6-glucosidase, amylosucrase, sucrose phosphorylase, and amylomaltase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QRW14378.1|GH13_22|GT5 | 0.0 | 31 | 2323 | 23 | 2353 |
| QRV85802.1|GH13_22|GT5 | 0.0 | 31 | 2323 | 23 | 2353 |
| QRV99867.1|GH13_22|GT5 | 0.0 | 31 | 2323 | 23 | 2353 |
| AAW44814.1|GH13_22|GT5|2.4.1.183 | 0.0 | 15 | 2323 | 16 | 2430 |
| AAR97368.1|GH13_22|GT5|2.4.1.183 | 0.0 | 15 | 2323 | 6 | 2420 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5A2B_A | 6.60e-12 | 63 | 190 | 28 | 146 | Crystal Structure of Anoxybacillus Alpha-amylase Provides Insights into a New Glycosyl Hydrolase Subclass [Anoxybacillus ayderensis],5A2C_A Crystal Structure of Anoxybacillus Alpha-amylase Provides Insights into a New Glycosyl Hydrolase Subclass [Anoxybacillus ayderensis] |
| 5A2A_A | 1.73e-11 | 75 | 190 | 6 | 112 | Crystal Structure of Anoxybacillus Alpha-amylase Provides Insights into a New Glycosyl Hydrolase Subclass [Anoxybacillus ayderensis] |
| 6SAO_A | 4.99e-11 | 73 | 190 | 4 | 109 | Structural and functional characterisation of three novel fungal amylases with enhanced stability and pH tolerance [Thamnidium elegans] |
| 4E2O_A | 5.29e-11 | 75 | 190 | 7 | 113 | Crystal structure of alpha-amylase from Geobacillus thermoleovorans, GTA, complexed with acarbose [Geobacillus thermoleovorans CCB_US3_UF5] |
| 3VM7_A | 5.98e-11 | 71 | 229 | 23 | 184 | Chain A, Alpha-amylase [Malbranchea cinnamomea] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| sp|Q9Y719|MOK13_SCHPO | 0.0 | 17 | 2323 | 6 | 2358 | Cell wall alpha-1,3-glucan synthase mok13 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=mok13 PE=3 SV=2 |
| sp|Q09854|MOK11_SCHPO | 0.0 | 31 | 2323 | 24 | 2397 | Cell wall alpha-1,3-glucan synthase mok11 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=mok11 PE=3 SV=2 |
| sp|Q9USK8|AGS1_SCHPO | 0.0 | 13 | 2323 | 7 | 2410 | Cell wall alpha-1,3-glucan synthase ags1 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=ags1 PE=1 SV=3 |
| sp|Q9UUL4|MOK12_SCHPO | 0.0 | 39 | 2321 | 32 | 2351 | Cell wall alpha-1,3-glucan synthase mok12 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=mok12 PE=3 SV=1 |
| sp|Q9Y704|MOK14_SCHPO | 1.10e-250 | 1035 | 2323 | 148 | 1369 | Cell wall alpha-1,3-glucan synthase mok14 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=mok14 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
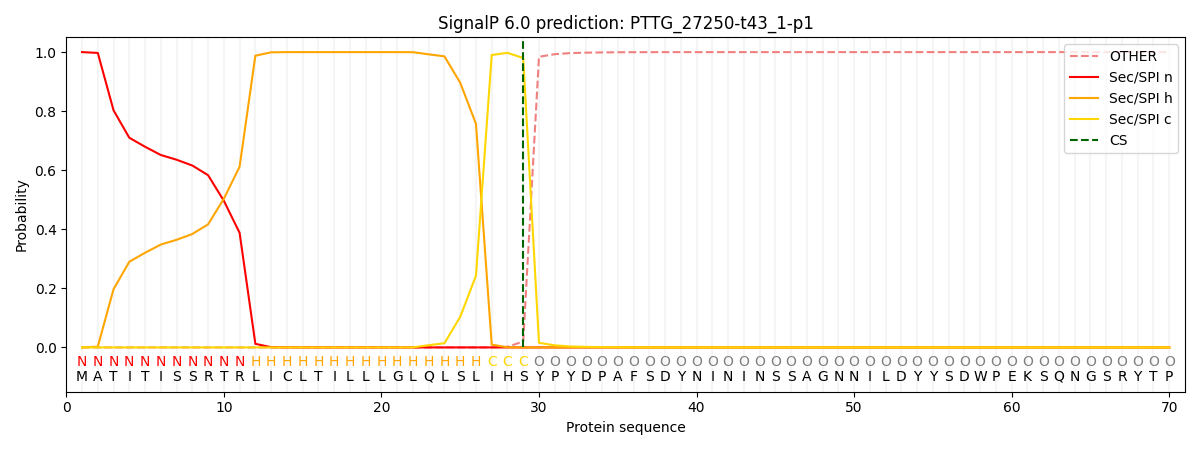
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000222 | 0.999729 | CS pos: 29-30. Pr: 0.9799 |
TMHMM Annotations download full data without filtering help
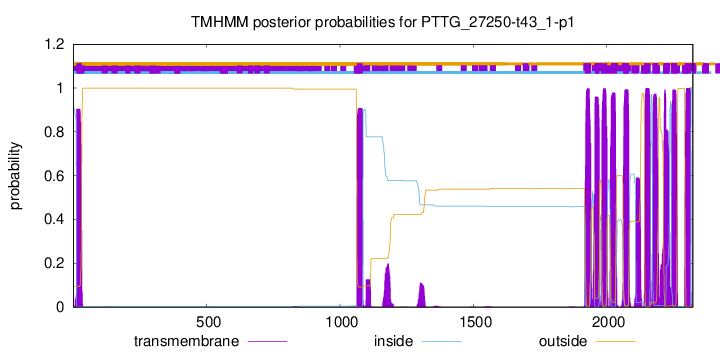
| Start | End |
|---|---|
| 12 | 31 |
| 1063 | 1085 |
| 1918 | 1940 |
| 1955 | 1972 |
| 1979 | 2001 |
| 2016 | 2038 |
| 2061 | 2083 |
| 2142 | 2164 |
| 2171 | 2193 |
| 2213 | 2235 |
| 2242 | 2264 |
| 2293 | 2315 |
