You are browsing environment: FUNGIDB
CAZyme Information: PTTG_00590-t43_1-p1
You are here: Home > Sequence: PTTG_00590-t43_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Puccinia triticina | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Basidiomycota; Pucciniomycetes; ; Pucciniaceae; Puccinia; Puccinia triticina | |||||||||||
| CAZyme ID | PTTG_00590-t43_1-p1 | |||||||||||
| CAZy Family | AA5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.4:20 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 22 | 301 | 8.6e-79 | 0.9822695035460993 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395098 | Cellulase | 1.29e-41 | 20 | 295 | 6 | 272 | Cellulase (glycosyl hydrolase family 5). |
| 225344 | BglC | 5.35e-05 | 21 | 301 | 49 | 370 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.33e-75 | 22 | 330 | 29 | 333 | |
| 4.71e-75 | 22 | 330 | 9 | 313 | |
| 6.26e-70 | 22 | 318 | 81 | 379 | |
| 8.83e-70 | 22 | 318 | 81 | 379 | |
| 1.19e-69 | 22 | 306 | 68 | 352 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.60e-60 | 22 | 318 | 8 | 317 | Chain A, Endoglucanase EG-II [Trichoderma reesei],3QR3_B Chain B, Endoglucanase EG-II [Trichoderma reesei] |
|
| 1.07e-53 | 22 | 303 | 27 | 320 | Structrue of a lucidum protein [Ganoderma lucidum],5D8W_B Structrue of a lucidum protein [Ganoderma lucidum],5D8Z_A Structrue of a lucidum protein [Ganoderma lucidum],5D8Z_B Structrue of a lucidum protein [Ganoderma lucidum] |
|
| 7.11e-47 | 22 | 297 | 27 | 304 | Structure of XacCel5A crystallized in the space group P41212 [Xanthomonas citri pv. citri str. 306] |
|
| 1.24e-46 | 22 | 297 | 27 | 304 | Crystal structure of XacCel5A in the native form [Xanthomonas citri pv. citri str. 306],4W7V_A Crystal structure of XacCel5A in complex with cellobiose [Xanthomonas citri pv. citri str. 306],4W7W_A High-resolution structure of XacCel5A in complex with cellopentaose [Xanthomonas citri pv. citri str. 306] |
|
| 3.86e-46 | 22 | 297 | 27 | 304 | Crystal structure of the endo-beta-1,4-glucanase (Xac0030) from Xanthomonas axonopodis pv. citri with the triple mutation His174Trp, Tyr211Ala and Lys227Arg. [Xanthomonas citri pv. citri str. 306],5HNN_B Crystal structure of the endo-beta-1,4-glucanase (Xac0030) from Xanthomonas axonopodis pv. citri with the triple mutation His174Trp, Tyr211Ala and Lys227Arg. [Xanthomonas citri pv. citri str. 306],5HNN_C Crystal structure of the endo-beta-1,4-glucanase (Xac0030) from Xanthomonas axonopodis pv. citri with the triple mutation His174Trp, Tyr211Ala and Lys227Arg. [Xanthomonas citri pv. citri str. 306] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.57e-70 | 22 | 318 | 81 | 379 | Manganese dependent endoglucanase Eg5A OS=Phanerodontia chrysosporium OX=2822231 GN=Eg5A PE=1 SV=2 |
|
| 1.60e-61 | 22 | 306 | 38 | 322 | Endoglucanase 1 OS=Saitozyma flava OX=5416 GN=CMC1 PE=2 SV=1 |
|
| 1.01e-58 | 22 | 318 | 98 | 407 | Endoglucanase EG-II OS=Hypocrea jecorina OX=51453 GN=egl2 PE=1 SV=1 |
|
| 7.75e-58 | 22 | 318 | 98 | 407 | Endoglucanase EG-II OS=Hypocrea jecorina (strain ATCC 56765 / BCRC 32924 / NRRL 11460 / Rut C-30) OX=1344414 GN=egl2 PE=1 SV=1 |
|
| 1.11e-39 | 9 | 290 | 107 | 390 | Endoglucanase OS=Ralstonia solanacearum (strain GMI1000) OX=267608 GN=egl PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
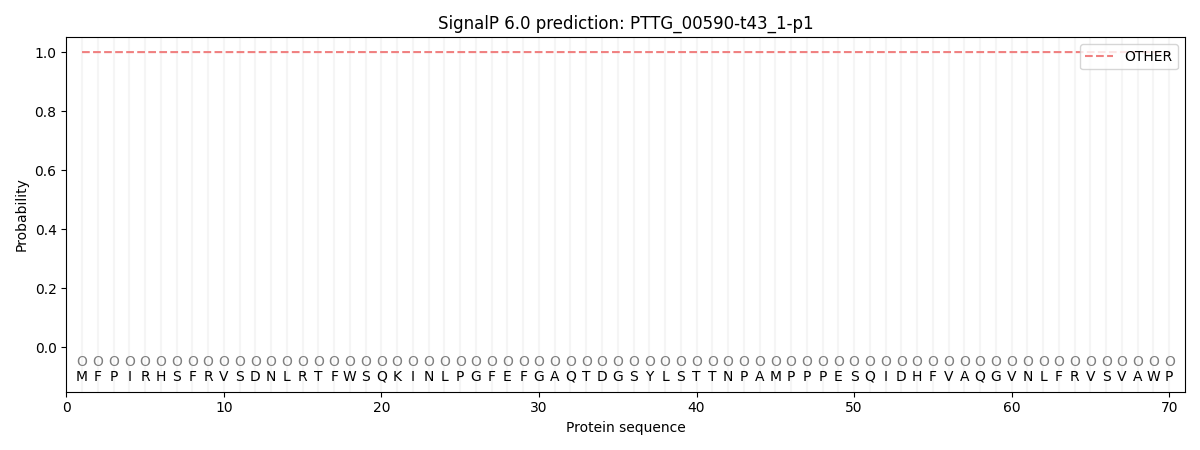
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000050 | 0.000010 |
