You are browsing environment: FUNGIDB
CAZyme Information: PSURA_76318T0-p1
You are here: Home > Sequence: PSURA_76318T0-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phytophthora ramorum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Peronosporaceae; Phytophthora; Phytophthora ramorum | |||||||||||
| CAZyme ID | PSURA_76318T0-p1 | |||||||||||
| CAZy Family | GH17 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH16 | 65 | 288 | 2.3e-62 | 0.6042944785276073 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 397841 | SKN1 | 1.44e-70 | 26 | 606 | 83 | 478 | Beta-glucan synthesis-associated protein (SKN1). This family consists of the beta-glucan synthesis-associated proteins KRE6 and SKN1. Beta1,6-Glucan is a key component of the yeast cell wall, interconnecting cell wall proteins, beta1,3-glucan, and chitin. It has been postulated that the synthesis of beta1,6-glucan begins in the endoplasmic reticulum with the formation of protein-bound primer structures and that these primer structures are extended in the Golgi complex by two putative glucosyltransferases that are functionally redundant, Kre6 and Skn1. This is followed by maturation steps at the cell surface and by coupling to other cell wall macromolecules. |
| 185689 | GH16_fungal_KRE6_glucanase | 5.11e-54 | 61 | 570 | 1 | 295 | Saccharomyces cerevisiae KRE6 and related glucanses, member of glycosyl hydrolase family 16. KRE6 is a Saccharomyces cerevisiae glucanase that participates in the synthesis of beta-1,6-glucan, a major structural component of the cell wall. It is a golgi membrane protein required for normal beta-1,6-glucan levels in the cell wall. KRE6 is closely realted to laminarinase, a glycosyl hydrolase family 16 member that hydrolyzes 1,3-beta-D-glucosidic linkages in 1,3-beta-D-glucans such as laminarins, curdlans, paramylons, and pachymans, with very limited action on mixed-link (1,3-1,4-)-beta-D-glucans. |
| 398415 | ELMO_CED12 | 3.74e-49 | 1724 | 1890 | 1 | 165 | ELMO/CED-12 family. This family represents a conserved domain which is found in a number of eukaryotic proteins including CED-12, ELMO I and ELMO II. ELMO1 is a component of signalling pathways that regulate phagocytosis and cell migration and is the mammalian orthologue of the C. elegans gene, ced-12. CED-12 is required for the engulfment of dying cells and cell migration. In mammalian cells, ELMO1 interacts with Dock180 as part of the CrkII/Dock180/Rac pathway responsible for phagocytosis and cell migration. ELMO1 is ubiquitously expressed, although its expression is highest in the spleen, an organ rich in immune cells. ELMO1 has a PH domain and a polyproline sequence motif at its C-terminus which are not present in this alignment. |
| 411345 | gliding_GltJ | 1.79e-11 | 1347 | 1485 | 418 | 561 | adventurous gliding motility protein GltJ. Adventurous gliding motility protein GltJ, also known as AgmX, occurs in delta-proteobacteria such as Myxococcus xanthus. |
| 411345 | gliding_GltJ | 1.42e-10 | 1344 | 1463 | 423 | 538 | adventurous gliding motility protein GltJ. Adventurous gliding motility protein GltJ, also known as AgmX, occurs in delta-proteobacteria such as Myxococcus xanthus. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 2 | 1960 | 37 | 1147 | |
| 0.0 | 30 | 733 | 29 | 729 | |
| 0.0 | 26 | 698 | 26 | 699 | |
| 0.0 | 19 | 711 | 6 | 697 | |
| 3.62e-291 | 63 | 731 | 1 | 665 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.58e-07 | 1767 | 1908 | 352 | 496 | Crystal Structure of BAI1/ELMO2 complex [Homo sapiens],6IE1_A Crystal Structure of ELMO2(Engulfment and cell motility protein 2) [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.53e-40 | 29 | 646 | 295 | 715 | Beta-glucan synthesis-associated protein KRE6 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=KRE6 PE=1 SV=2 |
|
| 8.23e-39 | 29 | 650 | 347 | 771 | Beta-glucan synthesis-associated protein SKN1 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=SKN1 PE=1 SV=1 |
|
| 1.80e-35 | 30 | 621 | 325 | 731 | Beta-glucan synthesis-associated protein SKN1 OS=Candida albicans OX=5476 GN=SKN1 PE=2 SV=1 |
|
| 2.25e-34 | 40 | 623 | 219 | 622 | Uncharacterized beta-glucan synthesis-associated protein C23H3.11c OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=SPAC23H3.11c PE=3 SV=1 |
|
| 2.62e-31 | 30 | 623 | 321 | 736 | Beta-glucan synthesis-associated protein KRE6 OS=Candida albicans OX=5476 GN=KRE6 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
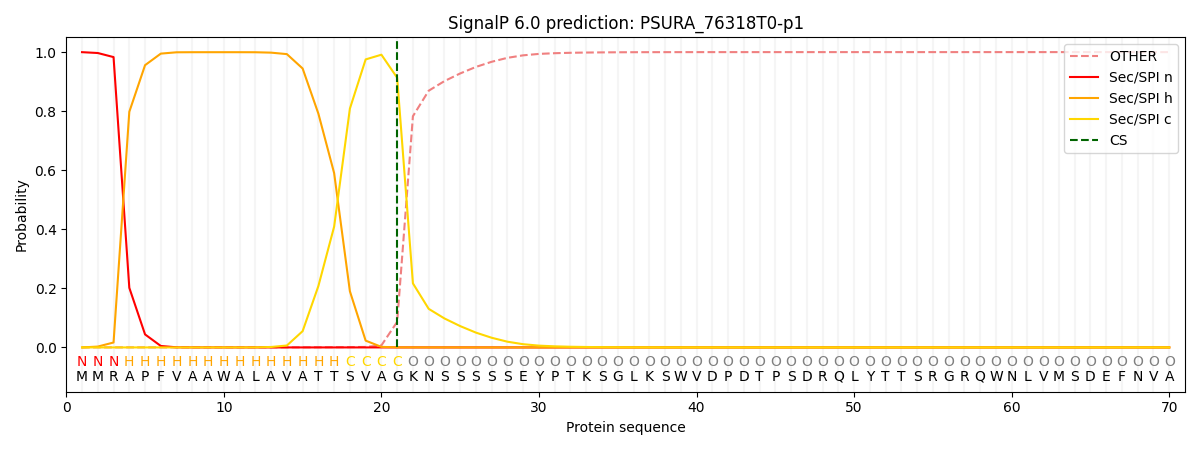
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.001479 | 0.998489 | CS pos: 21-22. Pr: 0.9132 |
