You are browsing environment: FUNGIDB
CAZyme Information: PSURA_39825T0-p1
You are here: Home > Sequence: PSURA_39825T0-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phytophthora ramorum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Peronosporaceae; Phytophthora; Phytophthora ramorum | |||||||||||
| CAZyme ID | PSURA_39825T0-p1 | |||||||||||
| CAZy Family | AA1 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL3 | 32 | 224 | 1.3e-80 | 0.9845360824742269 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 397360 | Pectate_lyase | 1.02e-90 | 26 | 228 | 2 | 198 | Pectate lyase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 2.63e-143 | 12 | 253 | 20 | 260 | |
| 2.51e-141 | 2 | 253 | 10 | 260 | |
| 4.58e-103 | 20 | 251 | 18 | 250 | |
| 2.65e-87 | 21 | 247 | 21 | 269 | |
| 2.65e-87 | 21 | 247 | 21 | 269 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.99e-27 | 38 | 184 | 6 | 147 | Crystal Structure Of Pectate Lyase From Bacillus Sp. Strain Ksm-P15. [Bacillus sp. KSM-P15] |
|
| 1.04e-21 | 43 | 176 | 16 | 143 | The liganded structure of C. bescii family 3 pectate lyase [Caldicellulosiruptor bescii DSM 6725],4EW9_B The liganded structure of C. bescii family 3 pectate lyase [Caldicellulosiruptor bescii DSM 6725] |
|
| 1.06e-21 | 43 | 176 | 17 | 144 | The crystal structure of family 3 pectate lyase from Caldicellulosiruptor bescii [Caldicellulosiruptor bescii],3T9G_B The crystal structure of family 3 pectate lyase from Caldicellulosiruptor bescii [Caldicellulosiruptor bescii] |
|
| 9.36e-21 | 43 | 176 | 25 | 152 | C. bescii Family 3 pectate lyase double mutant K108A in complex with trigalacturonic acid [Caldicellulosiruptor bescii DSM 6725],4Z03_B C. bescii Family 3 pectate lyase double mutant K108A in complex with trigalacturonic acid [Caldicellulosiruptor bescii DSM 6725] |
|
| 9.36e-21 | 43 | 176 | 25 | 152 | C. bescii Family 3 pectate lyase mutant E84A [Caldicellulosiruptor bescii DSM 6725],4Z05_B C. bescii Family 3 pectate lyase mutant E84A [Caldicellulosiruptor bescii DSM 6725] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.68e-66 | 29 | 246 | 28 | 239 | Probable pectate lyase G OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=plyG PE=3 SV=1 |
|
| 2.08e-63 | 29 | 246 | 43 | 257 | Probable pectate lyase D OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=plyD PE=3 SV=1 |
|
| 3.44e-62 | 29 | 247 | 23 | 236 | Probable pectate lyase D OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=plyD PE=3 SV=1 |
|
| 3.44e-62 | 29 | 247 | 23 | 236 | Probable pectate lyase D OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=plyD PE=3 SV=1 |
|
| 9.45e-62 | 29 | 247 | 22 | 235 | Probable pectate lyase D OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=plyD PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
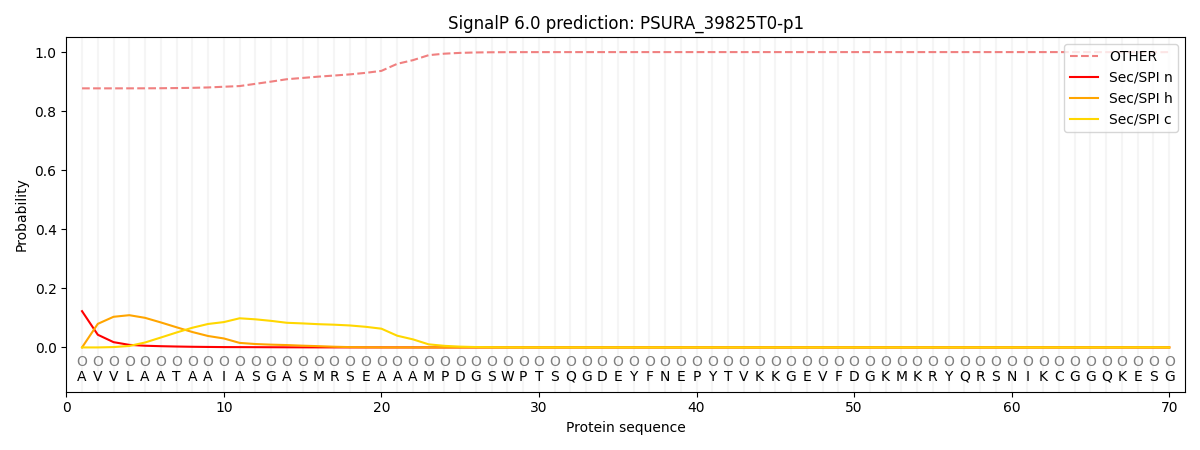
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.882676 | 0.117346 |
