You are browsing environment: FUNGIDB
CAZyme Information: PPTG_18414-t26_1-p1
You are here: Home > Sequence: PPTG_18414-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phytophthora parasitica | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Peronosporaceae; Phytophthora; Phytophthora parasitica | |||||||||||
| CAZyme ID | PPTG_18414-t26_1-p1 | |||||||||||
| CAZy Family | GT60 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.113:20 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH47 | 164 | 596 | 2.4e-134 | 0.9977578475336323 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396217 | Glyco_hydro_47 | 1.42e-139 | 164 | 596 | 1 | 453 | Glycosyl hydrolase family 47. Members of this family are alpha-mannosidases that catalyze the hydrolysis of the terminal 1,2-linked alpha-D-mannose residues in the oligo-mannose oligosaccharide Man(9)(GlcNAc)(2). |
| 240427 | PTZ00470 | 2.94e-83 | 148 | 599 | 63 | 521 | glycoside hydrolase family 47 protein; Provisional |
| 239047 | PA_GO-like | 3.77e-18 | 835 | 965 | 21 | 139 | PA_GO-like: Protease-associated domain containing proteins like Arabidopsis thaliana growth-on protein GRO10. This group contains various PA domain-containing proteins similar to the functionally uncharacterized Arabidopsis GRO10. The PA domain may be a protein-protein interaction domain. At peptidase active sites, the PA domain may participate in substrate binding and/or promoting conformational changes, which influence the stability and accessibility of the site to substrate. |
| 240122 | PA_subtilisin_1 | 4.90e-17 | 830 | 957 | 1 | 117 | PA_subtilisin_1: Protease-associated domain containing subtilisin-like proteases, subgroup 1. A subgroup of PA domain-containing subtilisin-like proteases. The significance of the PA domain to many of the proteins in which it is inserted is undetermined. It may be a protein-protein interaction domain. At peptidase active sites, the PA domain may participate in substrate binding and/or promoting conformational changes, which influence the stability and accessibility of the site to substrate. Proteins into which the PA domain is inserted include the following subtilisin-like proteases: i) melon cucumisin, ii) Arabidopsis thaliana Ara12, iii) Alnus glutinosa ag12, iv) members of the tomato P69 family, and v) tomato LeSBT2. However, these proteins belong to other subtilisin-like subgroups. Relatively little is known about proteins in this subgroup. |
| 238300 | PA | 7.79e-16 | 828 | 952 | 1 | 120 | PA: Protease-associated (PA) domain. The PA domain is an insert domain in a diverse fraction of proteases. The significance of the PA domain to many of the proteins in which it is inserted is undetermined. It may be a protein-protein interaction domain. At peptidase active sites, the PA domain may participate in substrate binding and/or promoting conformational changes, which influence the stability and accessibility of the site to substrate. Proteins into which the PA domain is inserted include the following: i) various signal peptide peptidases including, hSPPL2a and 2b which catalyze the intramembrane proteolysis of tumor necrosis factor alpha, ii) various proteins containing a C3H2C3 RING finger including, Arabidopsis ReMembR-H2 protein and various E3 ubiquitin ligases such as human GRAIL (gene related to anergy in lymphocytes), iii) EDEM3 (ER-degradation-enhancing mannosidase-like 3 protein), iv) various plant vacuolar sorting receptors such as Pisum sativum BP-80, v) glutamate carboxypeptidase II (GCPII), vi) yeast aminopeptidase Y, vii) Vibrio metschnikovii VapT, a sodium dodecyl sulfate (SDS) resistant extracellular alkaline serine protease, viii) lactocepin (a cell envelope-associated protease from Lactobacillus paracasei subsp. paracasei NCDO 151), ix) various subtilisin-like proteases such as melon Cucumisin, and x) human TfR (transferrin receptor) 1 and 2. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 76 | 966 | 1 | 967 | |
| 6.13e-249 | 1 | 936 | 43 | 878 | |
| 8.58e-246 | 6 | 936 | 33 | 870 | |
| 9.12e-130 | 152 | 598 | 30 | 470 | |
| 1.24e-129 | 152 | 598 | 30 | 470 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.16e-54 | 152 | 594 | 8 | 433 | Structure of The GH47 processing alpha-1,2-mannosidase from Caulobacter strain K31 [Caulobacter sp. K31],4AYP_A Structure of The GH47 processing alpha-1,2-mannosidase from Caulobacter strain K31 in complex with thiomannobioside [Caulobacter sp. K31],4AYQ_A Structure of The GH47 processing alpha-1,2-mannosidase from Caulobacter strain K31 in complex with mannoimidazole [Caulobacter sp. K31],4AYR_A Structure of The GH47 processing alpha-1,2-mannosidase from Caulobacter strain K31 in complex with noeuromycin [Caulobacter sp. K31],5MEH_A Crystal structure of alpha-1,2-mannosidase from Caulobacter K31 strain in complex with 1-deoxymannojirimycin [Caulobacter sp. K31],5NE5_A Crystal structure of family 47 alpha-1,2-mannosidase from Caulobacter K31 strain in complex with kifunensine [Caulobacter sp. K31] |
|
| 1.27e-52 | 159 | 596 | 7 | 451 | Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase and Man9GlcNAc2-PA complex [Homo sapiens] |
|
| 1.45e-52 | 159 | 596 | 12 | 456 | Crystal Structure Of Human Class I Alpha1,2-Mannosidase [Homo sapiens] |
|
| 1.52e-52 | 159 | 596 | 12 | 456 | Crystal Structure Of Human Class I Alpha1,2-Mannosidase In Complex With 1-Deoxymannojirimycin [Homo sapiens],1FO3_A Crystal Structure Of Human Class I Alpha1,2-Mannosidase In Complex With Kifunensine [Homo sapiens] |
|
| 1.85e-52 | 161 | 597 | 26 | 468 | Structure of mouse Golgi alpha-1,2-mannosidase IA and Man9GlcNAc2-PA complex [Mus musculus],5KKB_B Structure of mouse Golgi alpha-1,2-mannosidase IA and Man9GlcNAc2-PA complex [Mus musculus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.84e-126 | 149 | 624 | 32 | 519 | Alpha-mannosidase I MNS4 OS=Arabidopsis thaliana OX=3702 GN=MNS4 PE=1 SV=1 |
|
| 3.80e-119 | 133 | 597 | 13 | 479 | Alpha-mannosidase I MNS5 OS=Arabidopsis thaliana OX=3702 GN=MNS5 PE=1 SV=1 |
|
| 1.04e-114 | 161 | 594 | 39 | 480 | ER degradation-enhancing alpha-mannosidase-like protein 2 OS=Homo sapiens OX=9606 GN=EDEM2 PE=1 SV=2 |
|
| 1.42e-114 | 161 | 594 | 39 | 480 | ER degradation-enhancing alpha-mannosidase-like protein 2 OS=Mus musculus OX=10090 GN=Edem2 PE=1 SV=1 |
|
| 4.72e-101 | 154 | 596 | 122 | 581 | ER degradation-enhancing alpha-mannosidase-like protein 1 OS=Mus musculus OX=10090 GN=Edem1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
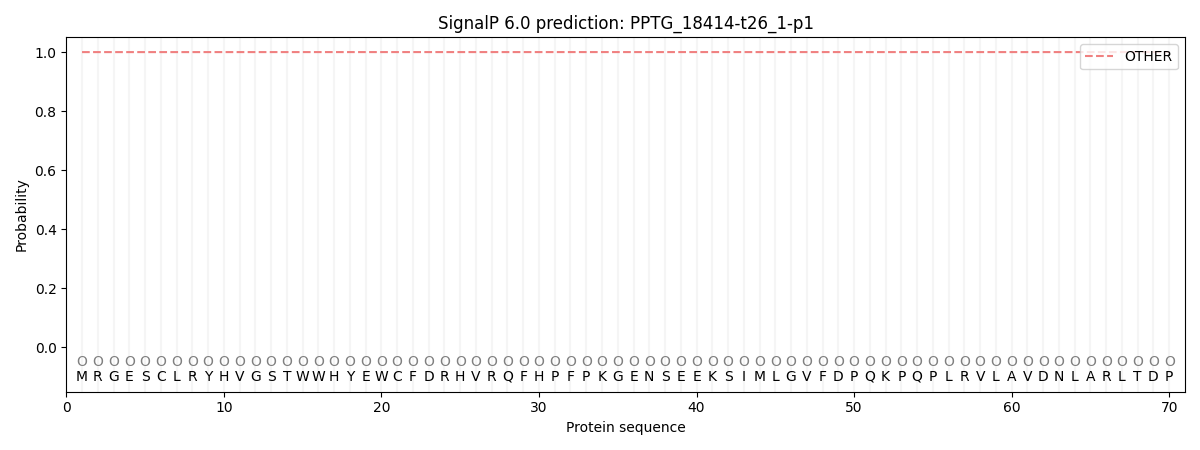
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000051 | 0.000000 |
