You are browsing environment: FUNGIDB
CAZyme Information: PPTG_07012-t26_1-p1
You are here: Home > Sequence: PPTG_07012-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phytophthora parasitica | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Peronosporaceae; Phytophthora; Phytophthora parasitica | |||||||||||
| CAZyme ID | PPTG_07012-t26_1-p1 | |||||||||||
| CAZy Family | GH140 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.176:83 | 3.2.1.132:6 | 3.2.1.4:2 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH7 | 26 | 458 | 3.9e-139 | 0.9927710843373494 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395677 | Glyco_hydro_7 | 0.0 | 26 | 457 | 1 | 434 | Glycosyl hydrolase family 7. |
| 153432 | GH7_CBH_EG | 2.85e-167 | 31 | 452 | 1 | 386 | Glycosyl hydrolase family 7. Glycosyl hydrolase family 7 contains eukaryotic endoglucanases (EGs) and cellobiohydrolases (CBHs) that hydrolyze glycosidic bonds using a double-displacement mechanism. This leads to a net retention of the conformation at the anomeric carbon. Both enzymes work synergistically in the degradation of cellulose,which is the main component of plant cell wall, and is composed of beta-1,4 linked glycosyl units. EG cleaves the beta-1,4 linkages of cellulose and CBH cleaves off cellobiose disaccharide units from the reducing end of the chain. In general, the O-glycosyl hydrolases are a widespread group of enzymes that hydrolyze the glycosidic bond between two or more carbohydrates, or between a carbohydrate and a non-carbohydrate moiety. A glycosyl hydrolase classification system based on sequence similarity has led to the definition of more than 95 different families inlcuding glycoside hydrolase family 7. |
| 408732 | Por_Secre_tail | 0.005 | 116 | 190 | 7 | 65 | Secretion system C-terminal sorting domain. Species that include Porphyromonas gingivalis, Fibrobacter succinogenes, Flavobacterium johnsoniae, Cytophaga hutchinsonii, Gramella forsetii, Prevotella intermedia, and Salinibacter ruber have on average twenty or more copies of this C-terminal domain, associated with sorting to the outer membrane and covalent modification. This domain targets proteins to type IX secretion systems and is secreted then cleaved off by a C-terminal signal peptidease. Based on similarity to other families it is likely that this domain adopts an immunoglobulin like fold. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.55e-177 | 22 | 458 | 23 | 464 | |
| 5.87e-143 | 16 | 458 | 10 | 453 | |
| 6.70e-143 | 10 | 458 | 5 | 457 | |
| 8.18e-143 | 10 | 457 | 83 | 536 | |
| 9.50e-143 | 10 | 458 | 5 | 457 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.30e-140 | 25 | 458 | 2 | 434 | Chain A, cellobiohydrolase I catalytic domain [Rasamsonia emersonii],3PFJ_A Chain A, Cellobiohydrolase 1 catalytic domain [Rasamsonia emersonii],3PFX_A Chain A, Cellobiohydrolase 1 catalytic domain [Rasamsonia emersonii],3PFZ_A Chain A, Cellobiohydrolase 1 catalytic domain [Rasamsonia emersonii],3PL3_A Chain A, Cellobiohydrolase 1 catalytic domain [Rasamsonia emersonii] |
|
| 4.90e-139 | 25 | 458 | 2 | 427 | Chain A, EXOGLUCANASE I [Phanerodontia chrysosporium],1H46_X Chain X, EXOGLUCANASE I [Phanerodontia chrysosporium],1Z3T_A Chain A, cellulase [Phanerodontia chrysosporium],1Z3V_A Chain A, cellulase [Phanerodontia chrysosporium],1Z3W_A Chain A, cellulase [Phanerodontia chrysosporium] |
|
| 3.47e-136 | 25 | 458 | 2 | 438 | Structure of Cellobiohydrolase 1 (Cel7A) from Heterobasidion annosum [Heterobasidion annosum],2YG1_A APO STRUCTURE OF CELLOBIOHYDROLASE 1 (CEL7A) FROM HETEROBASIDION ANNOSUM [Heterobasidion annosum],2YG1_B APO STRUCTURE OF CELLOBIOHYDROLASE 1 (CEL7A) FROM HETEROBASIDION ANNOSUM [Heterobasidion annosum] |
|
| 1.59e-134 | 25 | 458 | 2 | 437 | The 3-D structure of the cellobiohydrolase, Cel7A, from Aspergillus fumigatus [Aspergillus fumigatus],4V20_A The 3-D structure of the cellobiohydrolase, Cel7A, from Aspergillus fumigatus, disaccharide complex [Aspergillus fumigatus] |
|
| 7.64e-129 | 25 | 458 | 2 | 433 | The structure of P. funicolosum Cel7A [Talaromyces funiculosus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.43e-143 | 24 | 458 | 19 | 445 | Exoglucanase 1 OS=Phanerodontia chrysosporium OX=2822231 GN=CBH1 PE=2 SV=1 |
|
| 6.28e-139 | 15 | 458 | 9 | 450 | Probable 1,4-beta-D-glucan cellobiohydrolase A OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=cbhA PE=3 SV=1 |
|
| 6.28e-139 | 15 | 458 | 9 | 450 | Probable 1,4-beta-D-glucan cellobiohydrolase A OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=cbhA PE=3 SV=1 |
|
| 6.49e-139 | 15 | 458 | 9 | 450 | Probable 1,4-beta-D-glucan cellobiohydrolase A OS=Aspergillus clavatus (strain ATCC 1007 / CBS 513.65 / DSM 816 / NCTC 3887 / NRRL 1 / QM 1276 / 107) OX=344612 GN=cbhA PE=3 SV=1 |
|
| 2.04e-137 | 15 | 458 | 9 | 450 | Probable 1,4-beta-D-glucan cellobiohydrolase A OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=cbhA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
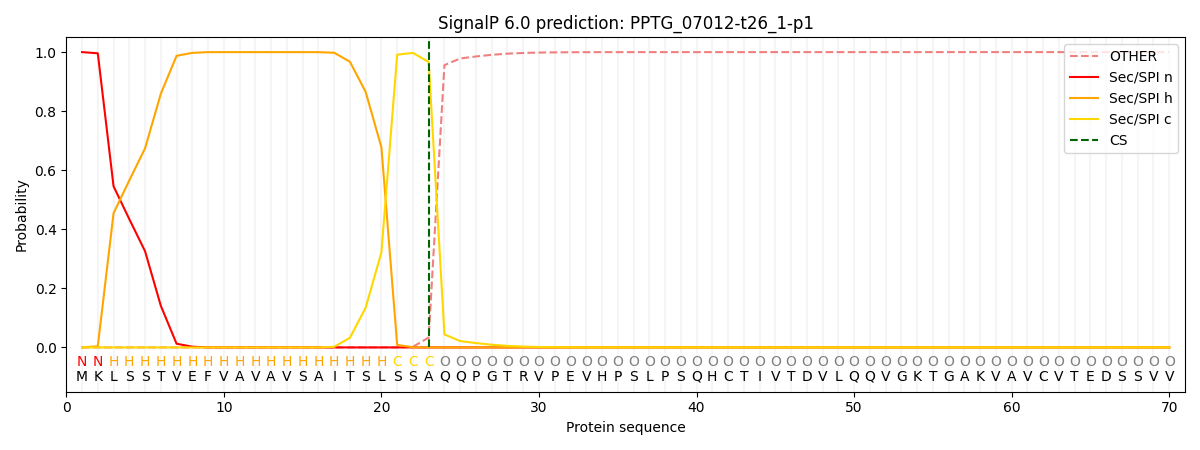
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000217 | 0.999735 | CS pos: 23-24. Pr: 0.9664 |
