You are browsing environment: FUNGIDB
CAZyme Information: PPTG_05922-t26_1-p1
You are here: Home > Sequence: PPTG_05922-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phytophthora parasitica | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Peronosporaceae; Phytophthora; Phytophthora parasitica | |||||||||||
| CAZyme ID | PPTG_05922-t26_1-p1 | |||||||||||
| CAZy Family | GH12 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH109 | 9 | 159 | 1.1e-20 | 0.39348370927318294 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 223745 | MviM | 4.77e-59 | 8 | 346 | 3 | 333 | Predicted dehydrogenase [General function prediction only]. |
| 396129 | GFO_IDH_MocA | 1.95e-23 | 9 | 129 | 1 | 119 | Oxidoreductase family, NAD-binding Rossmann fold. This family of enzymes utilize NADP or NAD. This family is called the GFO/IDH/MOCA family in swiss-prot. |
| 275173 | myo_inos_iolG | 3.13e-19 | 8 | 284 | 1 | 276 | inositol 2-dehydrogenase. All members of the seed alignment for this model are known or predicted inositol 2-dehydrogenase sequences co-clustered with other enzymes for catabolism of myo-inositol or closely related compounds. Inositol 2-dehydrogenase catalyzes the first step in inositol catabolism. Members of this family may vary somewhat in their ranges of acceptable substrates and some may act on analogs to myo-inositol rather than myo-inositol per se. [Energy metabolism, Sugars] |
| 182305 | PRK10206 | 1.72e-09 | 13 | 198 | 6 | 187 | putative oxidoreductase; Provisional |
| 183212 | PRK11579 | 5.32e-09 | 65 | 230 | 57 | 223 | putative oxidoreductase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.58e-10 | 61 | 254 | 82 | 291 | |
| 4.58e-10 | 61 | 254 | 82 | 291 | |
| 8.61e-10 | 64 | 158 | 124 | 218 | |
| 1.43e-09 | 9 | 158 | 61 | 217 | |
| 2.47e-09 | 9 | 158 | 45 | 201 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.73e-57 | 7 | 352 | 1 | 332 | Crystal structure of Mammalian Dimeric Dihydrodiol Dehydrogenase [Macaca fascicularis],2O4U_X Crystal structure of Mammalian Dimeric Dihydrodiol Dehydrogenase [Macaca fascicularis],2POQ_X Dimeric Dihydrodiol Dehydrogenase complexed with inhibitor, Isoascorbic acid [Macaca fascicularis],3OHS_X Crystal Structure of Mammalian Dimeric Dihydrodiol Dehydrogenase in complex with Dihydroxyacetone [Macaca fascicularis] |
|
| 1.15e-35 | 9 | 257 | 6 | 251 | Crystal Structure of an oxidoreductase from Enterococcus faecalis [Enterococcus faecalis],3E9M_B Crystal Structure of an oxidoreductase from Enterococcus faecalis [Enterococcus faecalis],3E9M_C Crystal Structure of an oxidoreductase from Enterococcus faecalis [Enterococcus faecalis],3E9M_D Crystal Structure of an oxidoreductase from Enterococcus faecalis [Enterococcus faecalis] |
|
| 2.18e-28 | 5 | 257 | 2 | 251 | CRYSTAL STRUCTURE OF putative oxidoreductase from Streptococcus agalactiae 2603V/r [Streptococcus agalactiae serogroup V] |
|
| 8.89e-24 | 6 | 225 | 21 | 239 | Crystal structure of probable oxidoreductase protein from Rhizobium etli CFN 42 [Rhizobium etli CFN 42],4HAD_B Crystal structure of probable oxidoreductase protein from Rhizobium etli CFN 42 [Rhizobium etli CFN 42],4HAD_C Crystal structure of probable oxidoreductase protein from Rhizobium etli CFN 42 [Rhizobium etli CFN 42],4HAD_D Crystal structure of probable oxidoreductase protein from Rhizobium etli CFN 42 [Rhizobium etli CFN 42] |
|
| 3.54e-21 | 8 | 200 | 27 | 216 | Crystal Structure of KijD10, a 3-ketoreductase from Actinomadura kijaniata incomplex with NADP [Actinomadura kijaniata],3RC1_A Crystal Structure of KijD10, a 3-ketoreductase from Actinomadura kijaniata incomplex with NADP and TDP-benzene [Actinomadura kijaniata],3RC2_A Crystal Structure of KijD10, a 3-ketoreductase from Actinomadura kijaniata in complex with TDP-benzene and NADP; open conformation [Actinomadura kijaniata] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.57e-67 | 7 | 353 | 1 | 333 | Trans-1,2-dihydrobenzene-1,2-diol dehydrogenase OS=Danio rerio OX=7955 GN=dhdh PE=2 SV=2 |
|
| 8.74e-63 | 10 | 349 | 4 | 329 | Trans-1,2-dihydrobenzene-1,2-diol dehydrogenase OS=Xenopus tropicalis OX=8364 GN=dhdh PE=2 SV=1 |
|
| 1.92e-61 | 10 | 349 | 4 | 329 | Trans-1,2-dihydrobenzene-1,2-diol dehydrogenase OS=Xenopus laevis OX=8355 GN=dhdh PE=2 SV=1 |
|
| 3.79e-59 | 7 | 352 | 1 | 333 | Trans-1,2-dihydrobenzene-1,2-diol dehydrogenase OS=Sus scrofa OX=9823 GN=DHDH PE=1 SV=1 |
|
| 1.06e-58 | 7 | 352 | 1 | 333 | Trans-1,2-dihydrobenzene-1,2-diol dehydrogenase OS=Bos taurus OX=9913 GN=DHDH PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
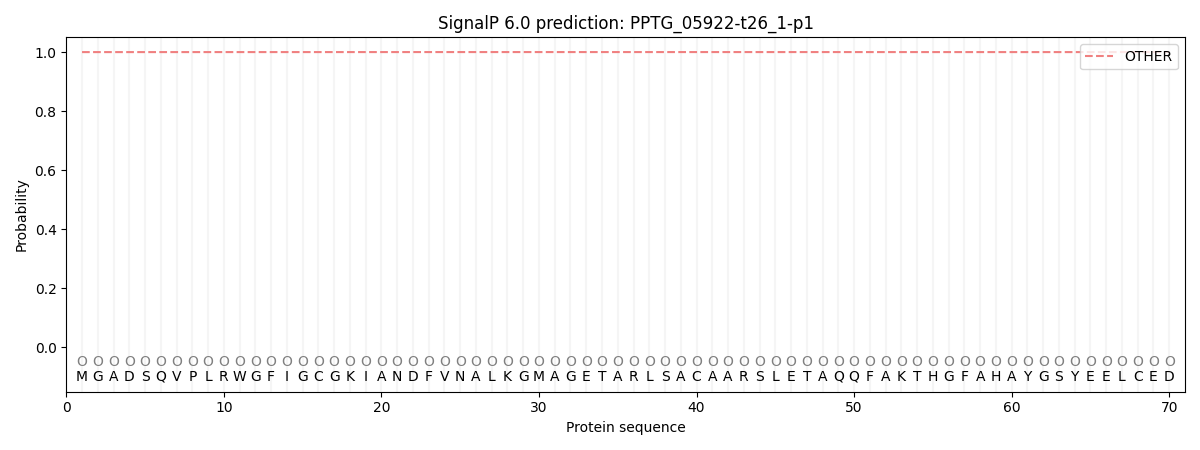
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000072 | 0.000000 |
