You are browsing environment: FUNGIDB
CAZyme Information: PNEJI1_003813-t26_1-p1
You are here: Home > Sequence: PNEJI1_003813-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Pneumocystis jirovecii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Pneumocystidomycetes; ; Pneumocystidaceae; Pneumocystis; Pneumocystis jirovecii | |||||||||||
| CAZyme ID | PNEJI1_003813-t26_1-p1 | |||||||||||
| CAZy Family | GT66 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 2.4.1.267:14 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT57 | 1 | 284 | 1.4e-69 | 0.6632016632016632 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 397324 | Alg6_Alg8 | 2.39e-73 | 1 | 285 | 148 | 461 | ALG6, ALG8 glycosyltransferase family. N-linked (asparagine-linked) glycosylation of proteins is mediated by a highly conserved pathway in eukaryotes, in which a lipid (dolichol phosphate)-linked oligosaccharide is assembled at the endoplasmic reticulum membrane prior to the transfer of the oligosaccharide moiety to the target asparagine residues. This oligosaccharide is composed of Glc(3)Man(9)GlcNAc(2). The addition of the three glucose residues is the final series of steps in the synthesis of the oligosaccharide precursor. Alg6 transfers the first glucose residue, and Alg8 transfers the second one. In the human alg6 gene, a C->T transition, which causes Ala333 to be replaced with Val, has been identified as the cause of a congenital disorder of glycosylation, designated as type Ic OMIM:603147. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.44e-120 | 1 | 285 | 206 | 466 | |
| 1.22e-83 | 1 | 285 | 404 | 696 | |
| 3.82e-81 | 1 | 285 | 190 | 471 | |
| 3.82e-81 | 1 | 285 | 190 | 471 | |
| 1.46e-80 | 1 | 284 | 234 | 544 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.33e-72 | 1 | 284 | 221 | 532 | Cryo-EM structure of yeast ALG6 in complex with 6AG9 Fab and Dol25-P-Glc [Saccharomyces cerevisiae],6SNI_X Cryo-EM structure of nanodisc reconstituted yeast ALG6 in complex with 6AG9 Fab [Saccharomyces cerevisiae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.79e-82 | 1 | 285 | 190 | 471 | Probable dolichyl pyrophosphate Man9GlcNAc2 alpha-1,3-glucosyltransferase OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=alg6 PE=3 SV=1 |
|
| 8.03e-72 | 1 | 284 | 203 | 514 | Dolichyl pyrophosphate Man9GlcNAc2 alpha-1,3-glucosyltransferase OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=ALG6 PE=1 SV=1 |
|
| 9.92e-39 | 2 | 280 | 171 | 470 | Dolichyl pyrophosphate Man9GlcNAc2 alpha-1,3-glucosyltransferase OS=Gallus gallus OX=9031 GN=ALG6 PE=2 SV=1 |
|
| 9.66e-38 | 2 | 285 | 210 | 498 | Probable dolichyl pyrophosphate Man9GlcNAc2 alpha-1,3-glucosyltransferase OS=Arabidopsis thaliana OX=3702 GN=At5g38460 PE=2 SV=1 |
|
| 7.92e-37 | 2 | 235 | 183 | 417 | Probable dolichyl pyrophosphate Man9GlcNAc2 alpha-1,3-glucosyltransferase OS=Dictyostelium discoideum OX=44689 GN=alg6 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
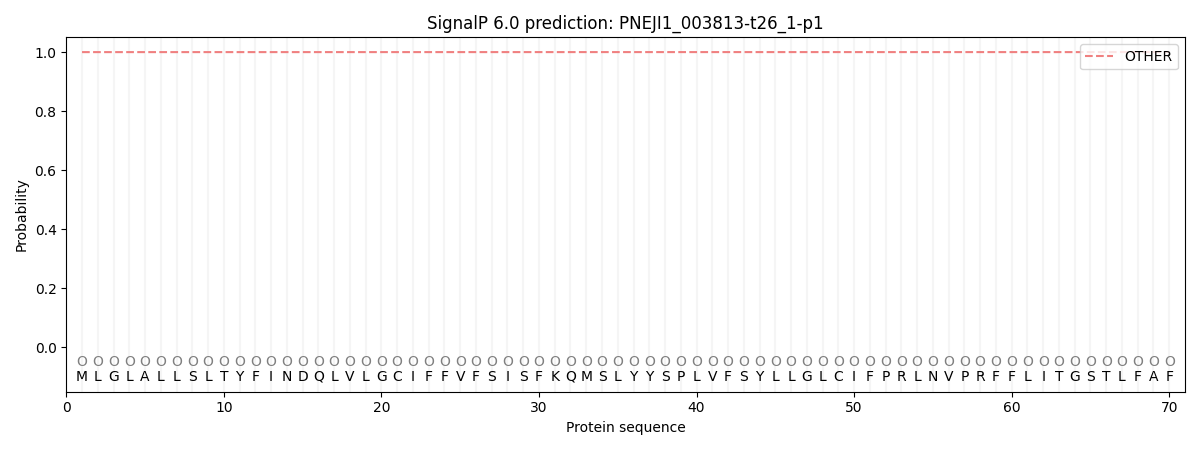
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000007 | 0.000000 |
TMHMM Annotations download full data without filtering help
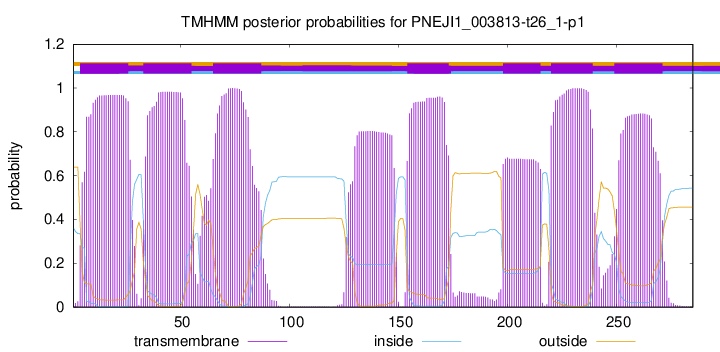
| Start | End |
|---|---|
| 4 | 26 |
| 33 | 55 |
| 65 | 87 |
| 154 | 173 |
| 198 | 215 |
| 220 | 239 |
| 249 | 271 |
