You are browsing environment: FUNGIDB
CAZyme Information: PIW_T001050-RA-p1
You are here: Home > Sequence: PIW_T001050-RA-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Globisporangium iwayamae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Pythiaceae; Globisporangium; Globisporangium iwayamae | |||||||||||
| CAZyme ID | PIW_T001050-RA-p1 | |||||||||||
| CAZy Family | AA17 | |||||||||||
| CAZyme Description | Alpha mannosyltransferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 2.4.1.122:14 | - |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT31 | 256 | 458 | 8.2e-39 | 0.984375 |
| GT10 | 787 | 951 | 1.7e-35 | 0.4438040345821326 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 250845 | Galactosyl_T | 1.15e-29 | 255 | 460 | 1 | 196 | Galactosyltransferase. This family includes the galactosyltransferases UDP-galactose:2-acetamido-2-deoxy-D-glucose3beta-galactosyltransferase and UDP-Gal:beta-GlcNAc beta 1,3-galactosyltranferase. Specific galactosyltransferases transfer galactose to GlcNAc terminal chains in the synthesis of the lacto-series oligosaccharides types 1 and 2. |
| 395683 | Glyco_transf_10 | 8.34e-15 | 814 | 938 | 17 | 130 | Glycosyltransferase family 10 (fucosyltransferase) C-term. This is the C-terminal domain of a family of fucosyltransferases. This enzyme transfers fucose from GDP-Fucose to GlcNAc in an alpha1,3 linkage. This family is known as glycosyltransferase family 10. The C-terminal domain is the likely binding-region for ADP (manuscript in publication). |
| 178735 | PLN03193 | 3.27e-11 | 241 | 459 | 140 | 348 | beta-1,3-galactosyltransferase; Provisional |
| 215596 | PLN03133 | 1.30e-10 | 239 | 449 | 384 | 579 | beta-1,3-galactosyltransferase; Provisional |
| 140237 | PTZ00210 | 5.95e-04 | 331 | 383 | 189 | 240 | UDP-GlcNAc-dependent glycosyltransferase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 2.48e-20 | 786 | 945 | 111 | 267 | |
| 3.22e-20 | 768 | 945 | 84 | 258 | |
| 1.25e-19 | 792 | 949 | 109 | 264 | |
| 5.04e-19 | 781 | 949 | 50 | 215 | |
| 9.46e-19 | 768 | 945 | 87 | 263 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.02e-14 | 794 | 959 | 174 | 328 | Crystal Structure of alpha1,3-Fucosyltransferase [Helicobacter pylori],2NZW_B Crystal Structure of alpha1,3-Fucosyltransferase [Helicobacter pylori],2NZW_C Crystal Structure of alpha1,3-Fucosyltransferase [Helicobacter pylori],2NZX_A Crystal Structure of alpha1,3-Fucosyltransferase with GDP [Helicobacter pylori],2NZX_B Crystal Structure of alpha1,3-Fucosyltransferase with GDP [Helicobacter pylori],2NZX_C Crystal Structure of alpha1,3-Fucosyltransferase with GDP [Helicobacter pylori],2NZY_A Crystal Structure of alpha1,3-Fucosyltransferase with GDP-fucose [Helicobacter pylori],2NZY_B Crystal Structure of alpha1,3-Fucosyltransferase with GDP-fucose [Helicobacter pylori],2NZY_C Crystal Structure of alpha1,3-Fucosyltransferase with GDP-fucose [Helicobacter pylori] |
|
| 5.42e-13 | 794 | 959 | 174 | 328 | Crystal Structure of alpha1,3-Fucosyltransferase [Helicobacter pylori],5ZOI_B Crystal Structure of alpha1,3-Fucosyltransferase [Helicobacter pylori] |
|
| 2.73e-10 | 230 | 480 | 99 | 339 | Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens],6WMN_B Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens],6WMN_C Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens],6WMN_D Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens],6WMO_A Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens],6WMO_B Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens] |
|
| 2.83e-10 | 230 | 480 | 105 | 345 | Chain A, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHK_B Chain B, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHK_C Chain C, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHK_D Chain D, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHL_A Chain A, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHL_B Chain B, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHM_A Chain A, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHM_B Chain B, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHN_A Chain A, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHO_A Chain A, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHO_B Chain B, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens] |
|
| 3.05e-10 | 230 | 480 | 119 | 359 | Chain A, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHI_B Chain B, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHI_C Chain C, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHI_D Chain D, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.39e-18 | 234 | 471 | 49 | 275 | Beta-1,3-galactosyltransferase 5 OS=Mus musculus OX=10090 GN=B3galt5 PE=2 SV=1 |
|
| 2.69e-18 | 234 | 471 | 51 | 277 | Beta-1,3-galactosyltransferase 5 (Fragment) OS=Pan troglodytes OX=9598 GN=B3GALT5 PE=3 SV=1 |
|
| 3.50e-18 | 781 | 949 | 405 | 570 | Putative fucosyltransferase R654 OS=Acanthamoeba polyphaga mimivirus OX=212035 GN=MIMI_R654 PE=3 SV=1 |
|
| 3.88e-18 | 234 | 471 | 51 | 277 | Beta-1,3-galactosyltransferase 5 (Fragment) OS=Pan paniscus OX=9597 GN=B3GALT5 PE=3 SV=1 |
|
| 2.22e-17 | 234 | 471 | 51 | 277 | Beta-1,3-galactosyltransferase 5 (Fragment) OS=Gorilla gorilla gorilla OX=9595 GN=B3GALT5 PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
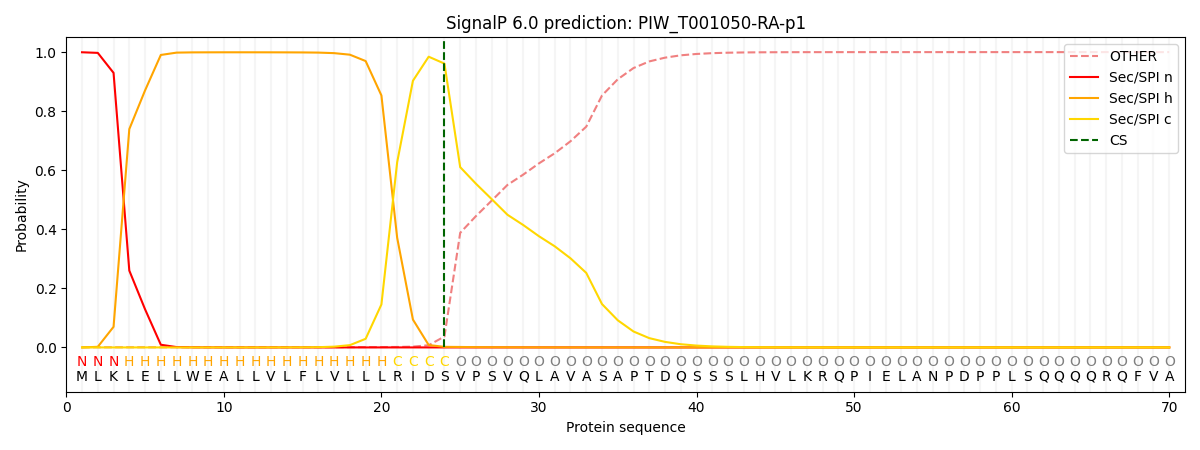
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.001335 | 0.998613 | CS pos: 24-25. Pr: 0.9611 |
