You are browsing environment: FUNGIDB
CAZyme Information: PHYSODRAFT_430630-t26_1-p1
You are here: Home > Sequence: PHYSODRAFT_430630-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phytophthora sojae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Peronosporaceae; Phytophthora; Phytophthora sojae | |||||||||||
| CAZyme ID | PHYSODRAFT_430630-t26_1-p1 | |||||||||||
| CAZy Family | GH30 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.15:10 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH28 | 16 | 332 | 5e-66 | 0.963076923076923 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395231 | Glyco_hydro_28 | 1.80e-75 | 20 | 332 | 1 | 321 | Glycosyl hydrolases family 28. Glycosyl hydrolase family 28 includes polygalacturonase EC:3.2.1.15 as well as rhamnogalacturonase A(RGase A), EC:3.2.1.-. These enzymes are important in cell wall metabolism. |
| 215540 | PLN03010 | 5.72e-25 | 72 | 302 | 141 | 357 | polygalacturonase |
| 178580 | PLN03003 | 4.39e-24 | 72 | 299 | 115 | 337 | Probable polygalacturonase At3g15720 |
| 215120 | PLN02188 | 2.10e-22 | 72 | 293 | 124 | 348 | polygalacturonase/glycoside hydrolase family protein |
| 177865 | PLN02218 | 3.60e-22 | 76 | 324 | 164 | 416 | polygalacturonase ADPG |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 2.52e-217 | 1 | 332 | 192 | 523 | |
| 5.40e-199 | 1 | 332 | 139 | 459 | |
| 5.41e-183 | 4 | 332 | 184 | 512 | |
| 4.13e-168 | 2 | 332 | 31 | 361 | |
| 3.78e-166 | 2 | 332 | 30 | 360 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.11e-76 | 3 | 332 | 1 | 336 | Structural insights into the processivity of endopolygalacturonase I from Aspergillus niger [Aspergillus niger],1NHC_B Structural insights into the processivity of endopolygalacturonase I from Aspergillus niger [Aspergillus niger],1NHC_C Structural insights into the processivity of endopolygalacturonase I from Aspergillus niger [Aspergillus niger],1NHC_D Structural insights into the processivity of endopolygalacturonase I from Aspergillus niger [Aspergillus niger],1NHC_E Structural insights into the processivity of endopolygalacturonase I from Aspergillus niger [Aspergillus niger],1NHC_F Structural insights into the processivity of endopolygalacturonase I from Aspergillus niger [Aspergillus niger] |
|
| 2.06e-74 | 4 | 332 | 10 | 344 | Chain A, Endo-polygalacturonase [Evansstolkia leycettana] |
|
| 2.06e-74 | 4 | 332 | 10 | 344 | Chain A, endo-polygalacturonase [Evansstolkia leycettana] |
|
| 2.30e-74 | 4 | 332 | 2 | 336 | Chain A, Endo-polygalacturonase [Evansstolkia leycettana] |
|
| 2.51e-74 | 4 | 328 | 2 | 333 | Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_B Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_C Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_D Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_E Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_F Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini],2IQ7_G Crystal structure of the polygalacturonase from Colletotrichum lupini and its implications for the interaction with polygalacturonase-inhibiting proteins [Colletotrichum lupini] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.73e-82 | 3 | 332 | 32 | 367 | Endopolygalacturonase B OS=Aspergillus flavus (strain ATCC MYA-384 / AF70) OX=1392242 GN=pgaB PE=2 SV=2 |
|
| 5.73e-82 | 3 | 332 | 32 | 367 | Probable endopolygalacturonase I OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=pgaI PE=3 SV=1 |
|
| 5.73e-82 | 3 | 332 | 32 | 367 | Endopolygalacturonase I OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=pgaI PE=1 SV=1 |
|
| 2.66e-80 | 2 | 332 | 32 | 368 | Probable endopolygalacturonase I OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=pgaI PE=3 SV=1 |
|
| 1.37e-78 | 4 | 328 | 26 | 355 | Polygalacturonase OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=PGU1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
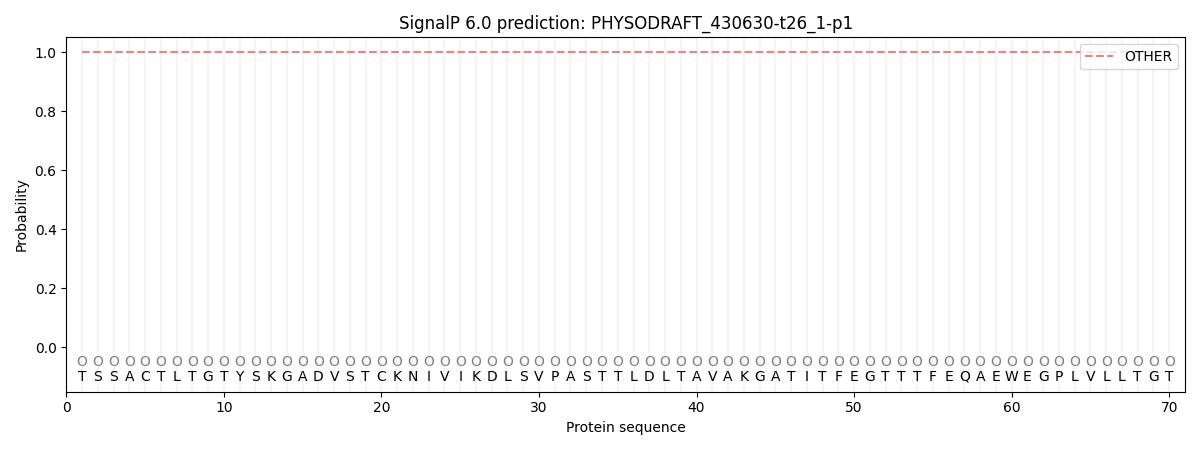
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000028 | 0.000001 |
