You are browsing environment: FUNGIDB
CAZyme Information: PHYSODRAFT_420719-t26_1-p1
You are here: Home > Sequence: PHYSODRAFT_420719-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phytophthora sojae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Peronosporaceae; Phytophthora; Phytophthora sojae | |||||||||||
| CAZyme ID | PHYSODRAFT_420719-t26_1-p1 | |||||||||||
| CAZy Family | GH30 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 2.4.1.-:43 | 2.4.1.149:17 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT31 | 23 | 225 | 1e-38 | 0.9791666666666666 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 250845 | Galactosyl_T | 1.27e-23 | 23 | 227 | 3 | 196 | Galactosyltransferase. This family includes the galactosyltransferases UDP-galactose:2-acetamido-2-deoxy-D-glucose3beta-galactosyltransferase and UDP-Gal:beta-GlcNAc beta 1,3-galactosyltranferase. Specific galactosyltransferases transfer galactose to GlcNAc terminal chains in the synthesis of the lacto-series oligosaccharides types 1 and 2. |
| 215596 | PLN03133 | 5.64e-13 | 8 | 222 | 387 | 585 | beta-1,3-galactosyltransferase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 5.49e-24 | 1 | 234 | 84 | 309 | |
| 7.73e-24 | 1 | 234 | 113 | 338 | |
| 7.73e-24 | 1 | 234 | 113 | 338 | |
| 7.73e-24 | 1 | 234 | 113 | 338 | |
| 7.73e-24 | 1 | 234 | 113 | 338 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.54e-15 | 7 | 234 | 110 | 325 | Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens],6WMN_B Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens],6WMN_C Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens],6WMN_D Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens],6WMO_A Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens],6WMO_B Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens] |
|
| 2.63e-15 | 7 | 234 | 116 | 331 | Chain A, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHK_B Chain B, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHK_C Chain C, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHK_D Chain D, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHL_A Chain A, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHL_B Chain B, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHM_A Chain A, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHM_B Chain B, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHN_A Chain A, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHO_A Chain A, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHO_B Chain B, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens] |
|
| 2.82e-15 | 7 | 234 | 130 | 345 | Chain A, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHI_B Chain B, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHI_C Chain C, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHI_D Chain D, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens] |
|
| 6.37e-15 | 7 | 234 | 110 | 325 | Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens],6WMM_B Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.37e-24 | 1 | 234 | 113 | 338 | UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 9 OS=Mus musculus OX=10090 GN=B3gnt9 PE=2 SV=1 |
|
| 8.95e-23 | 8 | 234 | 118 | 340 | UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 9 OS=Bos taurus OX=9913 GN=B3GNT9 PE=2 SV=1 |
|
| 3.22e-22 | 8 | 234 | 118 | 341 | UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 9 OS=Homo sapiens OX=9606 GN=B3GNT9 PE=2 SV=1 |
|
| 7.97e-22 | 5 | 255 | 129 | 366 | UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 7 OS=Rattus norvegicus OX=10116 GN=B3gnt7 PE=2 SV=1 |
|
| 5.50e-21 | 5 | 234 | 133 | 352 | UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 7 OS=Homo sapiens OX=9606 GN=B3GNT7 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
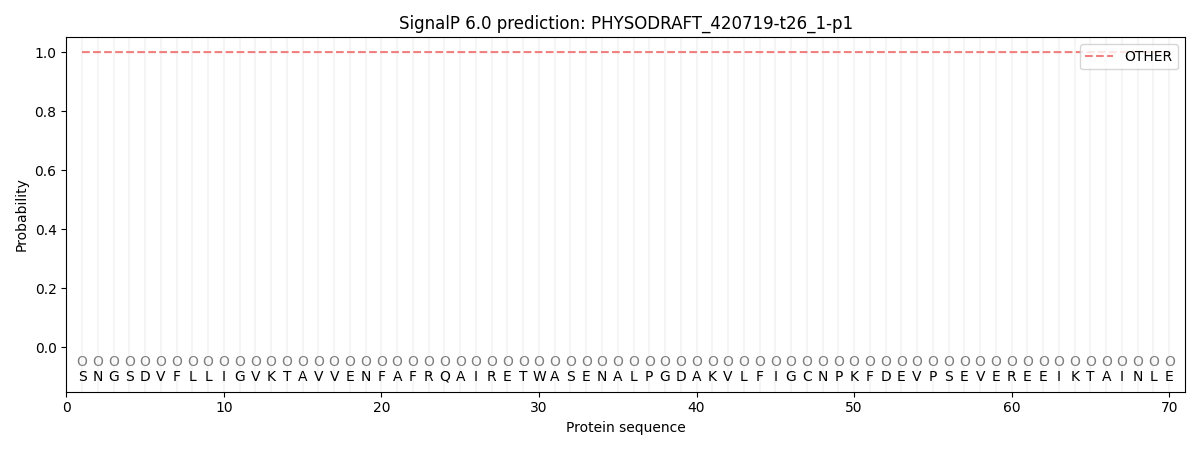
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000064 | 0.000000 |
