You are browsing environment: FUNGIDB
CAZyme Information: PHYSODRAFT_318573-t26_1-p1
You are here: Home > Sequence: PHYSODRAFT_318573-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phytophthora sojae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Peronosporaceae; Phytophthora; Phytophthora sojae | |||||||||||
| CAZyme ID | PHYSODRAFT_318573-t26_1-p1 | |||||||||||
| CAZy Family | CE8 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 2.4.1.122:14 |
|---|
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 250845 | Galactosyl_T | 7.23e-23 | 88 | 290 | 2 | 194 | Galactosyltransferase. This family includes the galactosyltransferases UDP-galactose:2-acetamido-2-deoxy-D-glucose3beta-galactosyltransferase and UDP-Gal:beta-GlcNAc beta 1,3-galactosyltranferase. Specific galactosyltransferases transfer galactose to GlcNAc terminal chains in the synthesis of the lacto-series oligosaccharides types 1 and 2. |
| 215596 | PLN03133 | 4.91e-13 | 72 | 285 | 385 | 583 | beta-1,3-galactosyltransferase; Provisional |
| 178735 | PLN03193 | 1.75e-06 | 74 | 286 | 141 | 345 | beta-1,3-galactosyltransferase; Provisional |
| 367085 | Fringe | 0.002 | 175 | 208 | 86 | 119 | Fringe-like. The drosophila protein fringe (FNG) is a glucosaminyltransferase that controls the response of the Notch receptor to specific ligands. FNG is localized to the Golgi apparatus (not secreted as previously thought). Modification of Notch occurs through glycosylation by FNG. The xenopus homolog, lunatic fringe, has been implicated in a variety of functions. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.76e-20 | 71 | 286 | 87 | 296 | |
| 3.28e-18 | 68 | 279 | 82 | 286 | |
| 3.28e-18 | 68 | 279 | 82 | 286 | |
| 3.28e-18 | 68 | 279 | 82 | 286 | |
| 3.28e-18 | 68 | 279 | 82 | 286 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.00e-09 | 74 | 252 | 111 | 279 | Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens],6WMM_B Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens] |
|
| 4.00e-09 | 74 | 252 | 111 | 279 | Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens],6WMN_B Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens],6WMN_C Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens],6WMN_D Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens],6WMO_A Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens],6WMO_B Human poly-N-acetyl-lactosamine synthase structure demonstrates a modular assembly of catalytic subsites for GT-A glycosyltransferases [Homo sapiens] |
|
| 4.08e-09 | 74 | 252 | 117 | 285 | Chain A, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHK_B Chain B, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHK_C Chain C, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHK_D Chain D, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHL_A Chain A, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHL_B Chain B, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHM_A Chain A, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHM_B Chain B, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHN_A Chain A, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHO_A Chain A, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHO_B Chain B, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens] |
|
| 4.26e-09 | 74 | 252 | 131 | 299 | Chain A, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHI_B Chain B, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHI_C Chain C, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens],7JHI_D Chain D, N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase 2 [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.34e-19 | 68 | 279 | 83 | 287 | Lactosylceramide 1,3-N-acetyl-beta-D-glucosaminyltransferase OS=Rattus norvegicus OX=10116 GN=B3gnt5 PE=2 SV=2 |
|
| 5.83e-19 | 68 | 279 | 82 | 286 | Lactosylceramide 1,3-N-acetyl-beta-D-glucosaminyltransferase OS=Mus musculus OX=10090 GN=B3gnt5 PE=2 SV=1 |
|
| 1.50e-18 | 71 | 279 | 87 | 289 | Lactosylceramide 1,3-N-acetyl-beta-D-glucosaminyltransferase A OS=Danio rerio OX=7955 GN=b3gnt5a PE=2 SV=1 |
|
| 2.85e-17 | 72 | 329 | 371 | 614 | Hydroxyproline O-galactosyltransferase GALT3 OS=Arabidopsis thaliana OX=3702 GN=GALT3 PE=2 SV=1 |
|
| 4.24e-17 | 68 | 279 | 84 | 288 | Lactosylceramide 1,3-N-acetyl-beta-D-glucosaminyltransferase OS=Homo sapiens OX=9606 GN=B3GNT5 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
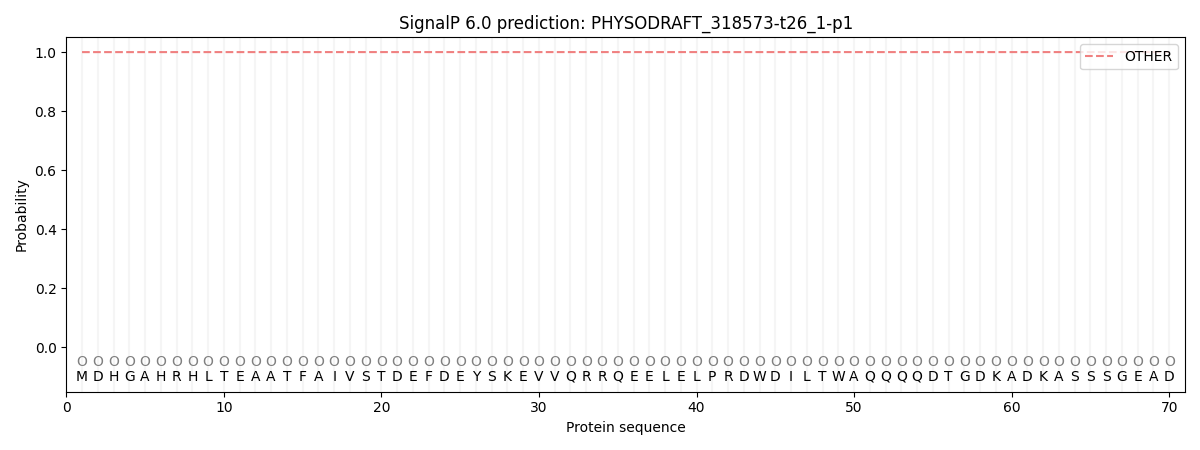
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000030 | 0.000001 |
