You are browsing environment: FUNGIDB
CAZyme Information: PHYSODRAFT_257416-t26_1-p1
You are here: Home > Sequence: PHYSODRAFT_257416-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phytophthora sojae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Peronosporaceae; Phytophthora; Phytophthora sojae | |||||||||||
| CAZyme ID | PHYSODRAFT_257416-t26_1-p1 | |||||||||||
| CAZy Family | AA17 | |||||||||||
| CAZyme Description | pectin methyl esterase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.1.1.11:1 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE8 | 36 | 311 | 1.9e-76 | 0.9236111111111112 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395871 | Pectinesterase | 5.17e-51 | 38 | 312 | 1 | 276 | Pectinesterase. |
| 178051 | PLN02432 | 1.73e-50 | 35 | 332 | 13 | 291 | putative pectinesterase |
| 178372 | PLN02773 | 6.05e-46 | 44 | 314 | 14 | 282 | pectinesterase |
| 215173 | PLN02304 | 6.27e-42 | 30 | 314 | 67 | 354 | probable pectinesterase |
| 215130 | PLN02217 | 1.09e-41 | 24 | 312 | 244 | 526 | probable pectinesterase/pectinesterase inhibitor |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 5.89e-176 | 1 | 339 | 1 | 338 | |
| 8.36e-176 | 1 | 339 | 1 | 338 | |
| 8.23e-159 | 13 | 339 | 15 | 344 | |
| 2.35e-158 | 13 | 339 | 15 | 344 | |
| 9.87e-143 | 1 | 339 | 1 | 343 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.33e-57 | 27 | 338 | 3 | 299 | Crystal Structure of the Pectin Methylesterase from Aspergillus niger in Deglycosylated Form [Aspergillus niger ATCC 1015] |
|
| 8.92e-57 | 27 | 338 | 3 | 299 | Crystal Structure of the Pectin Methylesterase from Aspergillus niger in Penultimately Deglycosylated Form (N-acetylglucosamine Stub at Asn84) [Aspergillus niger ATCC 1015] |
|
| 2.52e-33 | 34 | 312 | 8 | 283 | Pectin methylesterase from Carrot [Daucus carota] |
|
| 2.50e-30 | 45 | 312 | 13 | 279 | Chain A, Pectinesterase 1 [Solanum lycopersicum] |
|
| 1.48e-14 | 58 | 302 | 61 | 366 | The structure of rice weevil pectin methyl esterase [Sitophilus oryzae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.94e-78 | 13 | 339 | 6 | 330 | Pectinesterase OS=Aspergillus niger OX=5061 GN=pme1 PE=1 SV=1 |
|
| 3.17e-77 | 13 | 338 | 6 | 330 | Pectinesterase OS=Aspergillus aculeatus OX=5053 GN=pme1 PE=2 SV=1 |
|
| 2.04e-64 | 27 | 338 | 28 | 324 | Probable pectinesterase A OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=pmeA PE=3 SV=1 |
|
| 2.04e-64 | 27 | 338 | 28 | 324 | Probable pectinesterase A OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=pmeA PE=3 SV=1 |
|
| 1.14e-63 | 27 | 338 | 28 | 324 | Probable pectinesterase A OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=pmeA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
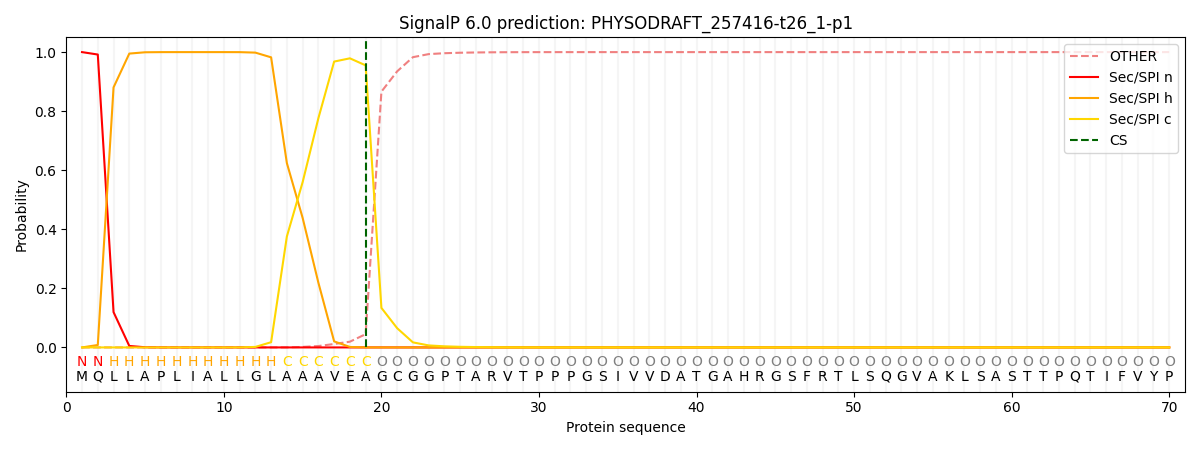
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000292 | 0.999677 | CS pos: 19-20. Pr: 0.9550 |
