You are browsing environment: FUNGIDB
CAZyme Information: PHYCI_87406T0-p1
You are here: Home > Sequence: PHYCI_87406T0-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
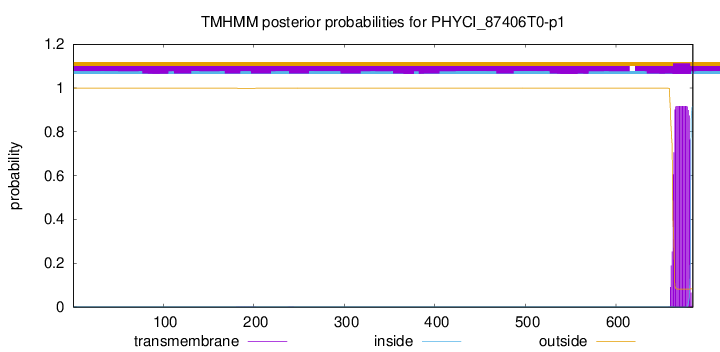
TMHMM annotations
Basic Information help
| Species | Phytophthora cinnamomi | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Peronosporaceae; Phytophthora; Phytophthora cinnamomi | |||||||||||
| CAZyme ID | PHYCI_87406T0-p1 | |||||||||||
| CAZy Family | GT30 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 19 | 518 | 3e-181 | 0.9981447124304267 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 408348 | Glyco_hydro_5_C | 1.10e-20 | 563 | 648 | 1 | 86 | Glycoside hydrolase family 5 C-terminal domain. This is the C-terminal domain of endo-glycoceramidase II (EGC), a membrane-associated family 5 glycosidase pfam00150. The C-terminal domain assumes a beta-sandwich fold, which resembles that of many carbohydrate-binding modules. |
| 395098 | Cellulase | 8.53e-05 | 64 | 133 | 20 | 88 | Cellulase (glycosyl hydrolase family 5). |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.47e-268 | 2 | 682 | 6 | 706 | |
| 7.49e-233 | 10 | 653 | 51 | 735 | |
| 5.10e-223 | 10 | 676 | 43 | 684 | |
| 3.48e-178 | 10 | 615 | 1 | 591 | |
| 4.68e-167 | 17 | 653 | 11 | 632 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.10e-118 | 16 | 618 | 27 | 725 | Chain A, Cytoplasmic protein [Cryptococcus neoformans var. grubii H99],7LPO_B Chain B, Cytoplasmic protein [Cryptococcus neoformans var. grubii H99],7LPO_C Chain C, Cytoplasmic protein [Cryptococcus neoformans var. grubii H99],7LPO_D Chain D, Cytoplasmic protein [Cryptococcus neoformans var. grubii H99],7LPP_A Chain A, Cytoplasmic protein [Cryptococcus neoformans var. grubii H99],7LPP_B Chain B, Cytoplasmic protein [Cryptococcus neoformans var. grubii H99],7LPP_C Chain C, Cytoplasmic protein [Cryptococcus neoformans var. grubii H99],7LPP_D Chain D, Cytoplasmic protein [Cryptococcus neoformans var. grubii H99],7LPQ_A Chain A, Cytoplasmic protein [Cryptococcus neoformans var. grubii H99],7LPQ_B Chain B, Cytoplasmic protein [Cryptococcus neoformans var. grubii H99],7LPQ_C Chain C, Cytoplasmic protein [Cryptococcus neoformans var. grubii H99],7LPQ_D Chain D, Cytoplasmic protein [Cryptococcus neoformans var. grubii H99] |
|
| 8.25e-10 | 2 | 253 | 34 | 226 | Chain A, Putative secreted endoglycosylceramidase [Rhodococcus hoagii 103S],5CCU_B Chain B, Putative secreted endoglycosylceramidase [Rhodococcus hoagii 103S] |
|
| 8.25e-10 | 2 | 253 | 34 | 226 | Chain A, Putative secreted endoglycosylceramidase [Rhodococcus hoagii 103S],5J14_B Chain B, Putative secreted endoglycosylceramidase [Rhodococcus hoagii 103S],5J7Z_A Chain A, Putative secreted endoglycosylceramidase [Rhodococcus hoagii 103S],5J7Z_B Chain B, Putative secreted endoglycosylceramidase [Rhodococcus hoagii 103S] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.24e-128 | 17 | 653 | 22 | 662 | Glucosylceramidase OS=Rhizopus delemar (strain RA 99-880 / ATCC MYA-4621 / FGSC 9543 / NRRL 43880) OX=246409 GN=ERC1 PE=1 SV=1 |
|
| 4.51e-122 | 17 | 539 | 13 | 569 | Glucosylceramidase OS=Neosartorya fumigata OX=746128 GN=egc1 PE=1 SV=1 |
|
| 5.34e-117 | 14 | 658 | 10 | 753 | Ergosteryl-beta-glucosidase OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=EGH1 PE=1 SV=1 |
|
| 4.16e-95 | 10 | 605 | 16 | 635 | Glucosylceramidase OS=Cryptococcus neoformans var. grubii serotype A (strain H99 / ATCC 208821 / CBS 10515 / FGSC 9487) OX=235443 GN=EGC1 PE=1 SV=1 |
|
| 5.11e-08 | 16 | 253 | 42 | 218 | Endoglycoceramidase I OS=Rhodococcus hoagii (strain 103S) OX=685727 GN=REQ_38260 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
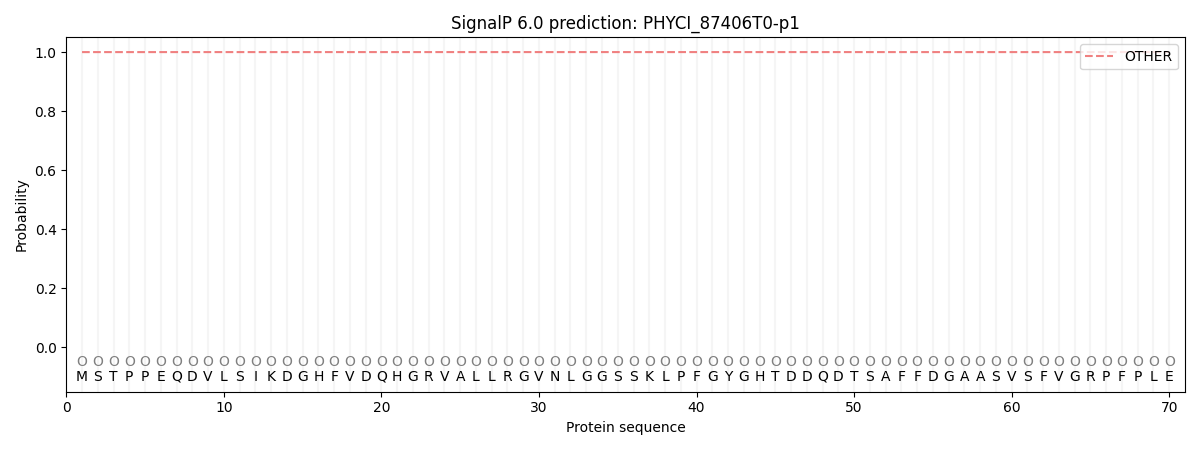
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000049 | 0.000000 |