You are browsing environment: FUNGIDB
CAZyme Information: PHYCI_533337T0-p1
You are here: Home > Sequence: PHYCI_533337T0-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
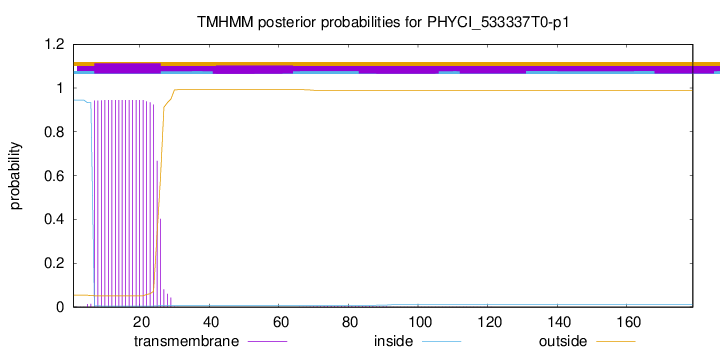
TMHMM annotations
Basic Information help
| Species | Phytophthora cinnamomi | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Peronosporaceae; Phytophthora; Phytophthora cinnamomi | |||||||||||
| CAZyme ID | PHYCI_533337T0-p1 | |||||||||||
| CAZy Family | GH7 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH53 | 26 | 170 | 6.7e-35 | 0.391812865497076 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 311610 | Glyco_hydro_53 | 1.62e-33 | 26 | 171 | 2 | 130 | Glycosyl hydrolase family 53. This domain belongs to family 53 of the glycosyl hydrolase classification. These enzymes are enzymes are endo-1,4- beta-galactanases (EC:3.2.1.89). The structure of this domain is known and has a TIM barrel fold. |
| 226385 | GanB | 8.25e-22 | 26 | 170 | 41 | 175 | Arabinogalactan endo-1,4-beta-galactosidase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.86e-27 | 14 | 171 | 6 | 144 | |
| 4.86e-27 | 14 | 171 | 6 | 144 | |
| 4.86e-27 | 14 | 171 | 6 | 144 | |
| 1.44e-26 | 26 | 171 | 39 | 174 | |
| 1.99e-26 | 26 | 171 | 21 | 146 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.62e-23 | 26 | 171 | 5 | 131 | Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Humicola insolens] |
|
| 3.22e-23 | 26 | 171 | 5 | 132 | Crystal Structure Of Beta-1,4-galactanase From Aspergillus Aculeatus At 293k [Aspergillus aculeatus],1FOB_A Crystal Structure Of Beta-1,4-galactanase From Aspergillus Aculeatus At 100k [Aspergillus aculeatus],6Q3R_A ASPERGILLUS ACULEATUS GALACTANASE [Aspergillus aculeatus],6Q3R_B ASPERGILLUS ACULEATUS GALACTANASE [Aspergillus aculeatus] |
|
| 2.84e-22 | 26 | 171 | 22 | 149 | Emericilla nidulans endo-beta-1,4-galactanase [Aspergillus nidulans] |
|
| 1.60e-20 | 26 | 171 | 5 | 131 | Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJS_B Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJS_C Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJS_D Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJU_A Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJU_B Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJU_C Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJU_D Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus] |
|
| 2.79e-11 | 22 | 165 | 5 | 133 | Beta-1,4-galactanase from Bacteroides thetaiotaomicron with galactose [Bacteroides thetaiotaomicron VPI-5482],6GPA_B Beta-1,4-galactanase from Bacteroides thetaiotaomicron with galactose [Bacteroides thetaiotaomicron VPI-5482] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.34e-26 | 26 | 171 | 21 | 146 | Probable arabinogalactan endo-beta-1,4-galactanase A OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=galA PE=3 SV=1 |
|
| 1.34e-26 | 26 | 171 | 21 | 146 | Probable arabinogalactan endo-beta-1,4-galactanase A OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=galA PE=3 SV=1 |
|
| 3.00e-25 | 1 | 171 | 1 | 153 | Probable arabinogalactan endo-beta-1,4-galactanase A OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=galA PE=3 SV=2 |
|
| 3.00e-25 | 1 | 171 | 1 | 153 | Probable arabinogalactan endo-beta-1,4-galactanase A OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=galA PE=3 SV=2 |
|
| 2.01e-24 | 26 | 171 | 21 | 148 | Probable arabinogalactan endo-beta-1,4-galactanase A OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=galA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
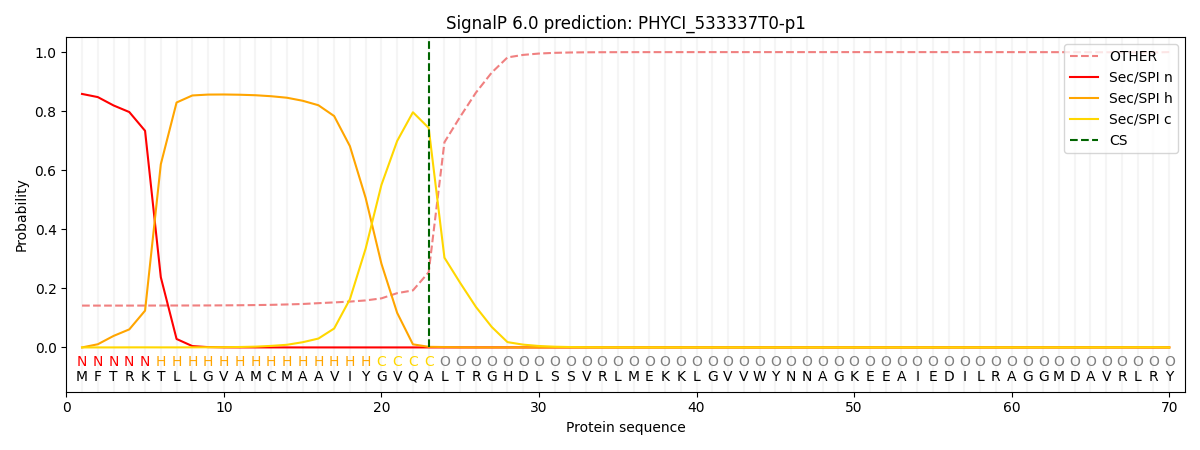
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.149802 | 0.850173 | CS pos: 23-24. Pr: 0.7433 |