You are browsing environment: FUNGIDB
CAZyme Information: PHYCI_32114T0-p1
You are here: Home > Sequence: PHYCI_32114T0-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phytophthora cinnamomi | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Peronosporaceae; Phytophthora; Phytophthora cinnamomi | |||||||||||
| CAZyme ID | PHYCI_32114T0-p1 | |||||||||||
| CAZy Family | PL3 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 239245 | TRX_family | 1.65e-07 | 9 | 92 | 1 | 86 | TRX family; composed of two groups: Group I, which includes proteins that exclusively encode a TRX domain; and Group II, which are composed of fusion proteins of TRX and additional domains. Group I TRX is a small ancient protein that alter the redox state of target proteins via the reversible oxidation of an active site dithiol, present in a CXXC motif, partially exposed at the protein's surface. TRX reduces protein disulfide bonds, resulting in a disulfide bond at its active site. Oxidized TRX is converted to the active form by TRX reductase, using reducing equivalents derived from either NADPH or ferredoxins. By altering their redox state, TRX regulates the functions of at least 30 target proteins, some of which are enzymes and transcription factors. It also plays an important role in the defense against oxidative stress by directly reducing hydrogen peroxide and certain radicals, and by serving as a reductant for peroxiredoxins. At least two major types of functional TRXs have been reported in most organisms; in eukaryotes, they are located in the cytoplasm and the mitochondria. Higher plants contain more types (at least 20 TRX genes have been detected in the genome of Arabidopsis thaliana), two of which (types f amd m) are located in the same compartment, the chloroplast. Also included in the alignment are TRX-like domains which show sequence homology to TRX but do not contain the redox active CXXC motif. Group II proteins, in addition to either a redox active TRX or a TRX-like domain, also contain additional domains, which may or may not possess homology to known proteins. |
| 395038 | Thioredoxin | 2.26e-06 | 2 | 80 | 1 | 83 | Thioredoxin. Thioredoxins are small enzymes that participate in redox reactions, via the reversible oxidation of an active centre disulfide bond. Some members with only the active site are not separated from the noise. |
| 239295 | PDI_a_PDIR | 2.71e-06 | 7 | 81 | 6 | 87 | PDIa family, PDIR subfamily; composed of proteins similar to human PDIR (for Protein Disulfide Isomerase Related). PDIR is composed of three redox active TRX (a) domains and an N-terminal redox inactive TRX-like (b) domain. Similar to PDI, it is involved in oxidative protein folding in the endoplasmic reticulum (ER) through its isomerase and chaperone activities. These activities are lower compared to PDI, probably due to PDIR acting only on a subset of proteins. PDIR is preferentially expressed in cells actively secreting proteins and its expression is induced by stress. Similar to PDI, the isomerase and chaperone activities of PDIR are independent; CXXC mutants lacking isomerase activity retain chaperone activity. |
| 273454 | pdi_dom | 4.17e-06 | 7 | 79 | 2 | 79 | protein disulfide-isomerase domain. This model describes a domain of eukaryotic protein disulfide isomerases, generally found in two copies. The high cutoff for total score reflects the expectation of finding both copies. The domain is similar to thioredoxin but the redox-active disulfide region motif is APWCGHCK. [Protein fate, Protein folding and stabilization] |
| 239303 | PDI_a_ERp46 | 5.59e-06 | 7 | 89 | 6 | 93 | PDIa family, endoplasmic reticulum protein 46 (ERp46) subfamily; ERp46 is an ER-resident protein containing three redox active TRX domains. Yeast complementation studies show that ERp46 can substitute for protein disulfide isomerase (PDI) function in vivo. It has been detected in many tissues, however, transcript and protein levels do not correlate in all tissues, suggesting regulation at a posttranscriptional level. An identical protein, named endoPDI, has been identified as an endothelial PDI that is highly expressed in the endothelium of tumors and hypoxic lesions. It has a protective effect on cells exposed to hypoxia. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.60e-106 | 219 | 751 | 56 | 571 | |
| 7.84e-106 | 219 | 751 | 80 | 595 | |
| 4.04e-98 | 219 | 753 | 57 | 564 | |
| 4.04e-98 | 219 | 753 | 57 | 564 | |
| 4.04e-98 | 219 | 753 | 57 | 564 |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
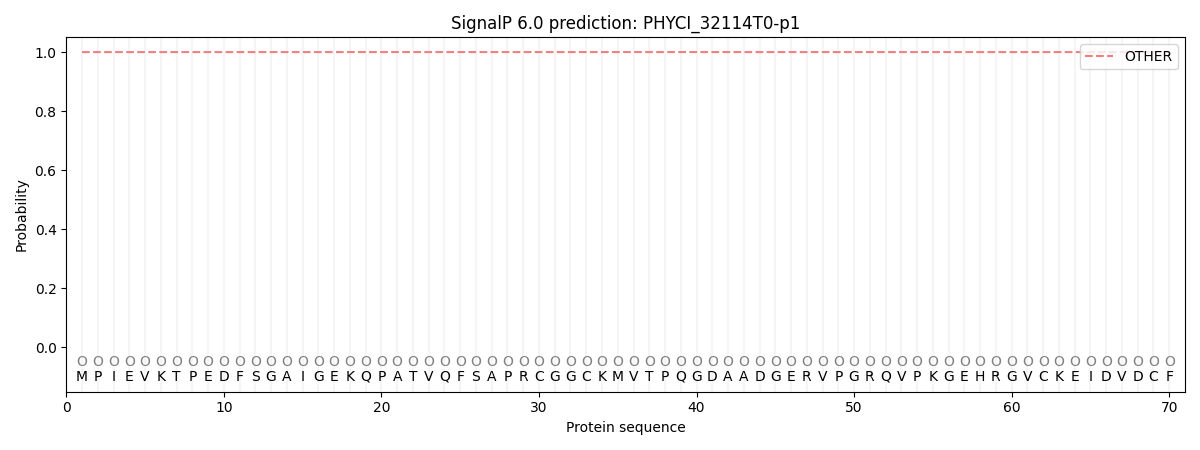
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000056 | 0.000002 |
