You are browsing environment: FUNGIDB
CAZyme Information: PHYCI_29977T0-p1
You are here: Home > Sequence: PHYCI_29977T0-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phytophthora cinnamomi | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Peronosporaceae; Phytophthora; Phytophthora cinnamomi | |||||||||||
| CAZyme ID | PHYCI_29977T0-p1 | |||||||||||
| CAZy Family | PL38 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 398135 | DUF455 | 2.22e-98 | 38 | 295 | 1 | 243 | Protein of unknown function (DUF455). |
| 225389 | COG2833 | 1.31e-54 | 35 | 299 | 11 | 264 | Uncharacterized conserved protein, contains ferritin-like DUF455 domain [Function unknown]. |
| 153097 | Ferritin_like | 5.81e-15 | 108 | 246 | 2 | 130 | Ferritin-like superfamily of diiron-containing four-helix-bundle proteins. Ferritin-like, diiron-carboxylate proteins participate in a range of functions including iron regulation, mono-oxygenation, and reactive radical production. These proteins are characterized by the fact that they catalyze dioxygen-dependent oxidation-hydroxylation reactions within diiron centers; one exception is manganese catalase, which catalyzes peroxide-dependent oxidation-reduction within a dimanganese center. Diiron-carboxylate proteins are further characterized by the presence of duplicate metal ligands, glutamates and histidines (ExxH) and two additional glutamates within a four-helix bundle. Outside of these conserved residues there is little obvious homology. Members include bacterioferritin, ferritin, rubrerythrin, aromatic and alkene monooxygenase hydroxylases (AAMH), ribonucleotide reductase R2 (RNRR2), acyl-ACP-desaturases (Acyl_ACP_Desat), manganese (Mn) catalases, demethoxyubiquinone hydroxylases (DMQH), DNA protecting proteins (DPS), and ubiquinol oxidases (AOX), and the aerobic cyclase system, Fe-containing subunit (ACSF). |
| 340831 | GT4_PimA-like | 2.34e-12 | 336 | 709 | 1 | 363 | phosphatidyl-myo-inositol mannosyltransferase. This family is most closely related to the GT4 family of glycosyltransferases and named after PimA in Propionibacterium freudenreichii, which is involved in the biosynthesis of phosphatidyl-myo-inositol mannosides (PIM) which are early precursors in the biosynthesis of lipomannans (LM) and lipoarabinomannans (LAM), and catalyzes the addition of a mannosyl residue from GDP-D-mannose (GDP-Man) to the position 2 of the carrier lipid phosphatidyl-myo-inositol (PI) to generate a phosphatidyl-myo-inositol bearing an alpha-1,2-linked mannose residue (PIM1). Glycosyltransferases catalyze the transfer of sugar moieties from activated donor molecules to specific acceptor molecules, forming glycosidic bonds. The acceptor molecule can be a lipid, a protein, a heterocyclic compound, or another carbohydrate residue. This group of glycosyltransferases is most closely related to the previously defined glycosyltransferase family 1 (GT1). The members of this family may transfer UDP, ADP, GDP, or CMP linked sugars. The diverse enzymatic activities among members of this family reflect a wide range of biological functions. The protein structure available for this family has the GTB topology, one of the two protein topologies observed for nucleotide-sugar-dependent glycosyltransferases. GTB proteins have distinct N- and C- terminal domains each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. The members of this family are found mainly in certain bacteria and archaea. |
| 404563 | Glyco_trans_1_4 | 2.89e-12 | 538 | 679 | 1 | 138 | Glycosyl transferases group 1. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 92 | 737 | 1 | 647 | |
| 1.09e-124 | 289 | 737 | 1 | 438 | |
| 6.14e-84 | 334 | 731 | 3 | 391 | |
| 6.02e-83 | 335 | 700 | 2 | 373 | |
| 1.61e-82 | 334 | 737 | 2 | 394 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.10e-08 | 780 | 855 | 575 | 648 | Crystal Structure of DesR, a beta-glucosidase from Streptomyces venezuelae in complex with D-glucose. [Streptomyces venezuelae],4I3G_B Crystal Structure of DesR, a beta-glucosidase from Streptomyces venezuelae in complex with D-glucose. [Streptomyces venezuelae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.72e-21 | 83 | 287 | 62 | 265 | Uncharacterized protein HI_0077 OS=Haemophilus influenzae (strain ATCC 51907 / DSM 11121 / KW20 / Rd) OX=71421 GN=HI_0077 PE=4 SV=1 |
|
| 1.27e-07 | 761 | 855 | 534 | 628 | Beta-glucosidase OS=Rhizobium radiobacter OX=358 GN=cbg-1 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
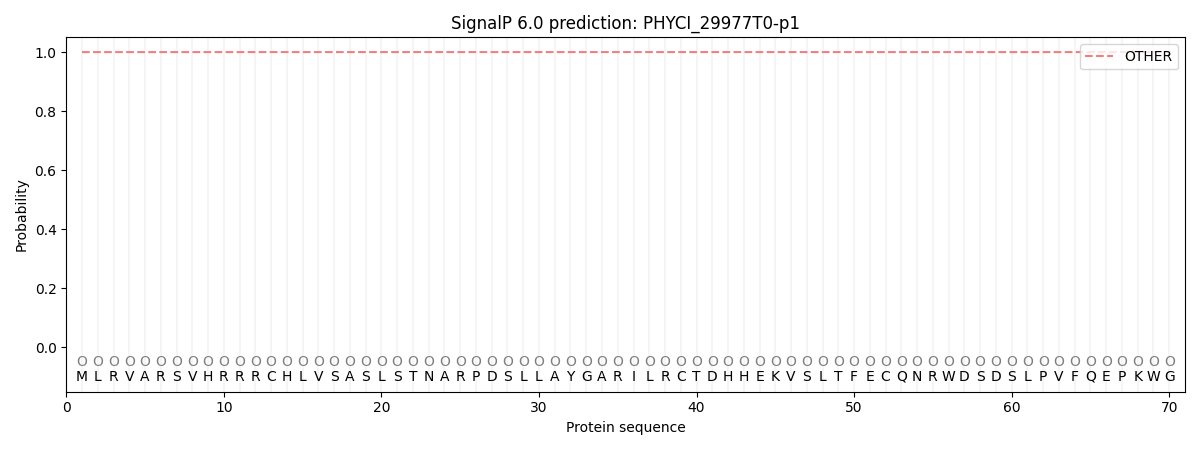
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000058 | 0.000002 |
