You are browsing environment: FUNGIDB
CAZyme Information: PHYCI_2956T0-p1
You are here: Home > Sequence: PHYCI_2956T0-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phytophthora cinnamomi | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Peronosporaceae; Phytophthora; Phytophthora cinnamomi | |||||||||||
| CAZyme ID | PHYCI_2956T0-p1 | |||||||||||
| CAZy Family | GH43 | |||||||||||
| CAZyme Description | Multicopper oxidases | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA1 | 56 | 532 | 2.8e-85 | 0.9357541899441341 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 274555 | ascorbase | 2.06e-69 | 62 | 541 | 39 | 526 | L-ascorbate oxidase, plant type. Members of this protein family are the copper-containing enzyme L-ascorbate oxidase (EC 1.10.3.3), also called ascorbase. This family is found in flowering plants, and shows greater sequence similarity to a family of laccases (EC 1.10.3.2) from plants than to other known ascorbate oxidases. |
| 177843 | PLN02191 | 5.01e-63 | 17 | 541 | 19 | 549 | L-ascorbate oxidase |
| 225043 | SufI | 1.39e-55 | 36 | 540 | 46 | 450 | Multicopper oxidase with three cupredoxin domains (includes cell division protein FtsP and spore coat protein CotA) [Cell cycle control, cell division, chromosome partitioning, Inorganic ion transport and metabolism, Cell wall/membrane/envelope biogenesis]. |
| 215324 | PLN02604 | 1.64e-55 | 62 | 538 | 62 | 546 | oxidoreductase |
| 274556 | laccase | 2.96e-45 | 62 | 537 | 41 | 518 | laccase, plant. Members of this protein family include the copper-containing enzyme laccase (EC 1.10.3.2), often several from a single plant species, and additional, uncharacterized, closely related plant proteins termed laccase-like multicopper oxidases. This protein family shows considerable sequence similarity to the L-ascorbate oxidase (EC 1.10.3.3) family. Laccases are enzymes of rather broad specificity, and classification of all proteins scoring about the trusted cutoff of this model as laccases may be appropriate. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 566 | 1 | 571 | |
| 0.0 | 9 | 566 | 11 | 569 | |
| 1.59e-199 | 17 | 563 | 21 | 547 | |
| 6.50e-187 | 7 | 563 | 2 | 539 | |
| 1.70e-79 | 15 | 560 | 34 | 553 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.72e-56 | 21 | 541 | 66 | 545 | Crystal structure of laccase from Botrytis aclada at 1.67 A resolution [Botrytis aclada],4X4K_A Structure of laccase from Botrytis aclada with full copper content [Botrytis aclada] |
|
| 9.97e-56 | 62 | 542 | 41 | 471 | Chain A, LACCASE 2 [Trametes versicolor] |
|
| 1.29e-55 | 21 | 541 | 66 | 545 | Structure of the L499M mutant of the laccase from B.aclada [Botrytis aclada] |
|
| 3.64e-55 | 62 | 542 | 42 | 471 | Single crystal serial study of the X-ray induced enzymatic reduction of molecular oxygen to water for laccase from Steccherinum murashkinskyi at sub-atomic resolution. Third structure of the series with 315 KGy dose. [Steccherinum murashkinskyi],6RHI_A Single crystal serial study of the X-ray induced enzymatic reduction of molecular oxygen to water for laccase from Steccherinum murashkinskyi at sub-atomic resolution. Ninth structure of the series with 1215 KGy dose. [Steccherinum murashkinskyi],6RHO_A Single crystal serial study of the X-ray induced enzymatic reduction of molecular oxygen to water for laccase from Steccherinum murashkinskyi at sub-atomic resolution. Twentieth structure of the series with 4065 KGy dose. [Steccherinum murashkinskyi],6RHP_A Single crystal serial study of the X-ray induced enzymatic reduction of molecular oxygen to water for laccase from Steccherinum murashkinskyi at sub-atomic resolution. Twenty first structure of the series with 4415 KGy dose (collected after refreezing). [Steccherinum murashkinskyi],6RHR_A Single crystal serial study of the inhibition of laccases from Steccherinum murashkinskyi by chloride anions at sub-atomic resolution. First structure of the series with 15 KGy dose. [Steccherinum murashkinskyi],6RHU_A Single crystal serial study of the inhibition of laccases from Steccherinum murashkinskyi by chloride anions at sub-atomic resolution. Second structure of the series with 165 KGy dose. [Steccherinum murashkinskyi],6RHX_A Single crystal serial study of the inhibition of laccases from Steccherinum murashkinskyi by chloride anions at sub-atomic resolution. Third structure of the series with 315 KGy dose. [Steccherinum murashkinskyi],6RI0_A Single crystal serial study of the inhibition of laccases from Steccherinum murashkinskyi by chloride anions at sub-atomic resolution. Ninth structure of the series with 1215 KGy dose. [Steccherinum murashkinskyi],6RI2_A Single crystal serial study of the inhibition of laccases from Steccherinum murashkinskyi by chloride anions at sub-atomic resolution. Twentieth structure of the series with 4065 KGy dose. [Steccherinum murashkinskyi],6RI4_A Single crystal serial study of the inhibition of laccases from Steccherinum murashkinskyi by fluoride anions at sub-atomic resolution. First structure of the series with 13 KGy dose. [Steccherinum murashkinskyi],6RI6_A Single crystal serial study of the inhibition of laccases from Steccherinum murashkinskyi by fluoride anions at sub-atomic resolution. Second structure of the series with 400 KGy dose. [Steccherinum murashkinskyi],6RI8_A Single crystal serial study of the inhibition of laccases from Steccherinum murashkinskyi by fluoride anions at sub-atomic resolution. Third structure of the series with 800 KGy dose. [Steccherinum murashkinskyi],6RII_A Single crystal serial study of the inhibition of laccases from Steccherinum murashkinskyi by fluoride anions at sub-atomic resolution. Fourth structure of the series with 1200 KGy dose. [Steccherinum murashkinskyi],6RIK_A Single crystal serial study of the inhibition of laccases from Steccherinum murashkinskyi by fluoride anions at sub-atomic resolution. Thirteenth structure of the series with 5200 KGy dose. [Steccherinum murashkinskyi],6RIL_A Single crystal serial study of the inhibition of laccases from Steccherinum murashkinskyi by fluoride anions at sub-atomic resolution. Fourteenth structure of the series with 5600 KGy dose (data was collected after refreezing). [Steccherinum murashkinskyi] |
|
| 3.71e-55 | 62 | 542 | 42 | 471 | Structural study of the X-ray induced enzymatic reduction of molecular oxygen to water for laccase from Steccherinum murashkinskyi. First structure of the series with 3 min total X-ray exposition time. [Steccherinum murashkinskyi],5MHU_A The study of the X-ray induced enzymatic reduction of molecular oxygen to water for laccase from Steccherinum murashkinskyi.The third structure of the series with total exposition time 63 min. [Steccherinum murashkinskyi],5MHV_A The study of the X-ray induced enzymatic reduction of molecular oxygen to water for laccase from Steccherinum murashkinskyi.The fourth structure of the series with total exposition time 93 min. [Steccherinum murashkinskyi] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.10e-69 | 14 | 541 | 53 | 524 | Laccase-1 OS=Botryotinia fuckeliana OX=40559 GN=lcc1 PE=2 SV=3 |
|
| 3.19e-62 | 57 | 538 | 106 | 551 | Oxidoreductase OpS5 OS=Beauveria bassiana (strain ARSEF 2860) OX=655819 GN=OpS5 PE=1 SV=1 |
|
| 7.18e-59 | 18 | 541 | 64 | 546 | Laccase-2 OS=Botryotinia fuckeliana OX=40559 GN=lcc2 PE=2 SV=1 |
|
| 1.70e-58 | 18 | 566 | 19 | 517 | Iron transport multicopper oxidase fetC OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=fetC PE=2 SV=1 |
|
| 2.76e-56 | 7 | 543 | 10 | 502 | Iron transport multicopper oxidase fio1 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=fio1 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
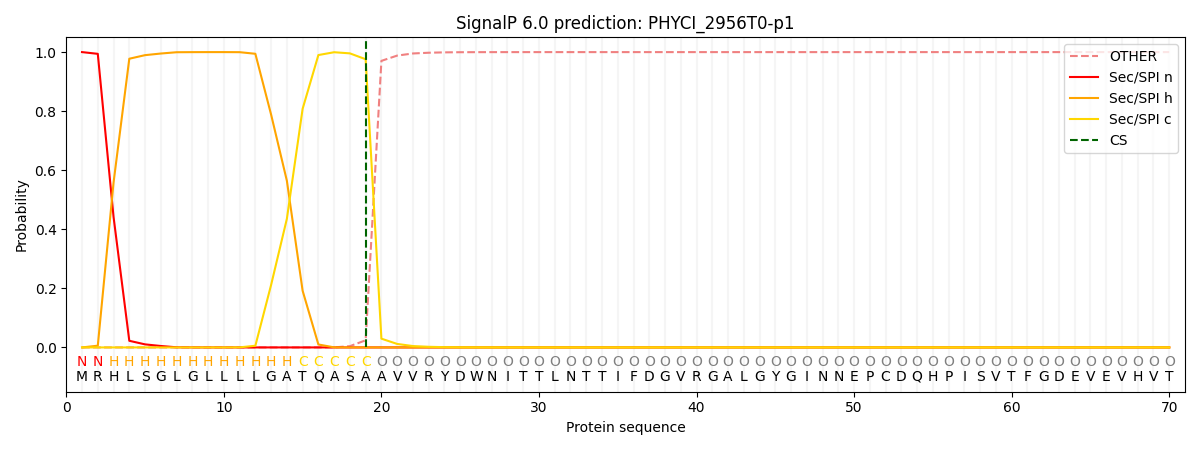
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000244 | 0.999738 | CS pos: 19-20. Pr: 0.9758 |
