You are browsing environment: FUNGIDB
CAZyme Information: PGTG_17514-t26_1-p1
You are here: Home > Sequence: PGTG_17514-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Puccinia graminis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Basidiomycota; Pucciniomycetes; ; Pucciniaceae; Puccinia; Puccinia graminis | |||||||||||
| CAZyme ID | PGTG_17514-t26_1-p1 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA1 | 58 | 590 | 3.8e-87 | 0.9776536312849162 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 132431 | ascorbOXfungal | 8.07e-159 | 42 | 597 | 9 | 532 | L-ascorbate oxidase, fungal type. This model describes a family of fungal ascorbate oxidases, within a larger family of multicopper oxidases that also includes plant ascorbate oxidases (TIGR03388), plant laccases and laccase-like proteins (TIGR03389), and related proteins. The member from Acremonium sp. HI-25 is characterized. |
| 274555 | ascorbase | 2.83e-96 | 54 | 610 | 14 | 537 | L-ascorbate oxidase, plant type. Members of this protein family are the copper-containing enzyme L-ascorbate oxidase (EC 1.10.3.3), also called ascorbase. This family is found in flowering plants, and shows greater sequence similarity to a family of laccases (EC 1.10.3.2) from plants than to other known ascorbate oxidases. |
| 259941 | CuRO_2_AAO_like_2 | 8.22e-89 | 174 | 335 | 1 | 161 | The second cupredoxin domain of plant Ascorbate oxidase homologs. This family includes plant laccases similar to ascorbate oxidase. Ascorbate oxidase catalyzes the oxidation of ascorbic acid to dehydroascorbic acid. It can detect levels of ascorbic acid and eliminate it. The biological function of ascorbate oxidase is still not clear. Ascorbate oxidase belongs to multicopper oxidase (MCO) family which couples oxidation of substrates with reduction of dioxygen to water. Although MCOs have diverse functions, majority of them have three cupredoxin domain repeats that include one mononuclear and one trinuclear copper center. The copper ions are bound in several sites: Type 1, Type 2, and/or Type 3. The ensemble of types 2 and 3 copper is called a trinuclear cluster. MCOs oxidize their substrate by accepting electrons at a mononuclear copper center and transferring them to the active site trinuclear copper center. The cupredoxin domain 2 of 3-domain MCOs has lost the ability to bind copper. |
| 177843 | PLN02191 | 2.61e-82 | 48 | 605 | 30 | 555 | L-ascorbate oxidase |
| 215324 | PLN02604 | 1.40e-76 | 54 | 605 | 37 | 555 | oxidoreductase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.04e-104 | 40 | 630 | 26 | 600 | |
| 3.02e-103 | 40 | 624 | 26 | 577 | |
| 3.02e-103 | 40 | 624 | 26 | 577 | |
| 3.02e-103 | 40 | 624 | 26 | 577 | |
| 4.78e-103 | 26 | 625 | 15 | 573 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.02e-73 | 56 | 605 | 18 | 532 | Refined Crystal Structure Of Ascorbate Oxidase At 1.9 Angstroms Resolution [Cucurbita pepo var. melopepo],1AOZ_B Refined Crystal Structure Of Ascorbate Oxidase At 1.9 Angstroms Resolution [Cucurbita pepo var. melopepo],1ASO_A X-Ray Structures And Mechanistic Implications Of Three Functional Derivatives Of Ascorbate Oxidase From Zucchini: Reduced-, Peroxide-, And Azide-Forms [Cucurbita pepo var. melopepo],1ASO_B X-Ray Structures And Mechanistic Implications Of Three Functional Derivatives Of Ascorbate Oxidase From Zucchini: Reduced-, Peroxide-, And Azide-Forms [Cucurbita pepo var. melopepo],1ASP_A X-ray Structures And Mechanistic Implications Of Three Functional Derivatives Of Ascorbate Oxidase From Zucchini: Reduced-, Peroxide-, And Azide-forms [Cucurbita pepo var. melopepo],1ASP_B X-ray Structures And Mechanistic Implications Of Three Functional Derivatives Of Ascorbate Oxidase From Zucchini: Reduced-, Peroxide-, And Azide-forms [Cucurbita pepo var. melopepo],1ASQ_A X-Ray Structures And Mechanistic Implications Of Three Functional Derivatives Of Ascorbate Oxidase From Zucchini: Reduced-, Peroxide-, And Azide-Forms [Cucurbita pepo var. melopepo],1ASQ_B X-Ray Structures And Mechanistic Implications Of Three Functional Derivatives Of Ascorbate Oxidase From Zucchini: Reduced-, Peroxide-, And Azide-Forms [Cucurbita pepo var. melopepo] |
|
| 1.39e-34 | 42 | 593 | 4 | 469 | Crystal structure of LacB from Trametes sp. AH28-2 [Trametes sp. AH28-2],3KW7_B Crystal structure of LacB from Trametes sp. AH28-2 [Trametes sp. AH28-2] |
|
| 6.52e-33 | 42 | 600 | 4 | 469 | Crystal Structure of Laccase from Cerrena sp. RSD1 [Cerrena],5Z1X_B Crystal Structure of Laccase from Cerrena sp. RSD1 [Cerrena],5Z22_A Crystal Structure of Laccase from Cerrena sp. RSD1 [Cerrena] |
|
| 1.87e-31 | 42 | 608 | 4 | 484 | Chain A, LACCASE 2 [Trametes versicolor] |
|
| 8.82e-31 | 60 | 594 | 22 | 477 | Crystal Structure of Fet3p, a Multicopper Oxidase that Functions in Iron Import [Saccharomyces cerevisiae],1ZPU_B Crystal Structure of Fet3p, a Multicopper Oxidase that Functions in Iron Import [Saccharomyces cerevisiae],1ZPU_C Crystal Structure of Fet3p, a Multicopper Oxidase that Functions in Iron Import [Saccharomyces cerevisiae],1ZPU_D Crystal Structure of Fet3p, a Multicopper Oxidase that Functions in Iron Import [Saccharomyces cerevisiae],1ZPU_E Crystal Structure of Fet3p, a Multicopper Oxidase that Functions in Iron Import [Saccharomyces cerevisiae],1ZPU_F Crystal Structure of Fet3p, a Multicopper Oxidase that Functions in Iron Import [Saccharomyces cerevisiae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.70e-100 | 42 | 632 | 31 | 636 | Laccase-like multicopper oxidase 1 OS=Myceliophthora thermophila (strain ATCC 42464 / BCRC 31852 / DSM 1799) OX=573729 GN=LMCO1 PE=1 SV=1 |
|
| 4.42e-99 | 42 | 623 | 28 | 591 | Multicopper oxidase aurL2 OS=Gibberella zeae (strain ATCC MYA-4620 / CBS 123657 / FGSC 9075 / NRRL 31084 / PH-1) OX=229533 GN=aurL2 PE=2 SV=1 |
|
| 4.90e-74 | 97 | 597 | 1 | 489 | Multicopper oxidase terE OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=terE PE=1 SV=1 |
|
| 6.98e-73 | 65 | 606 | 54 | 560 | L-ascorbate oxidase OS=Nicotiana tabacum OX=4097 GN=AAO PE=2 SV=1 |
|
| 9.96e-73 | 56 | 605 | 48 | 562 | L-ascorbate oxidase OS=Cucurbita maxima OX=3661 GN=AAO PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
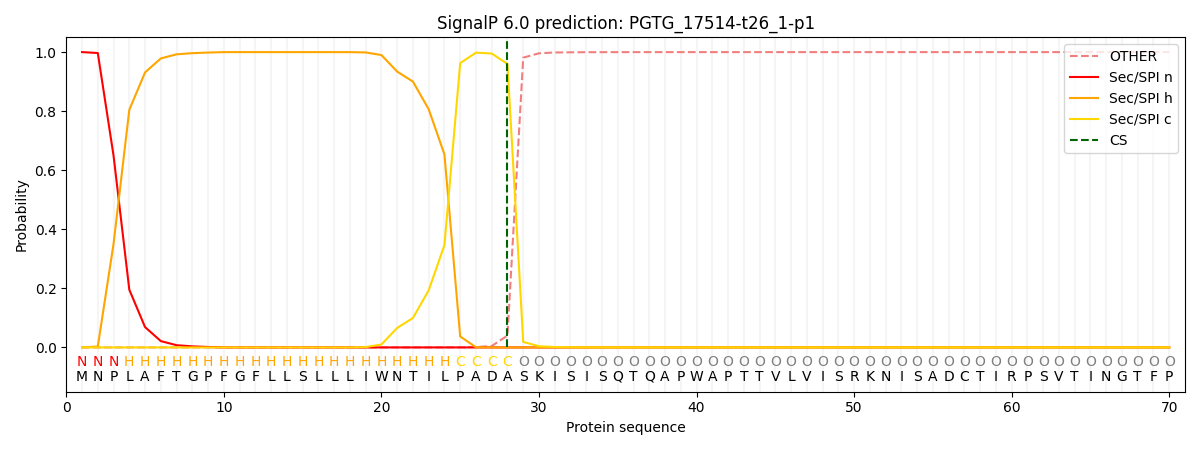
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000224 | 0.999743 | CS pos: 28-29. Pr: 0.9597 |
