You are browsing environment: FUNGIDB
CAZyme Information: PGTG_11725-t26_1-p1
You are here: Home > Sequence: PGTG_11725-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
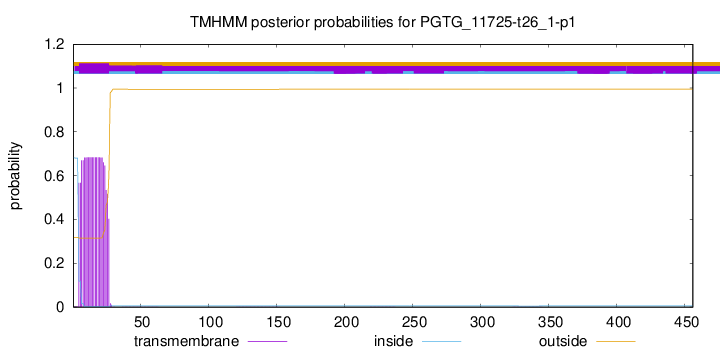
TMHMM annotations
Basic Information help
| Species | Puccinia graminis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Basidiomycota; Pucciniomycetes; ; Pucciniaceae; Puccinia; Puccinia graminis | |||||||||||
| CAZyme ID | PGTG_11725-t26_1-p1 | |||||||||||
| CAZy Family | GH38 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.4:20 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 47 | 346 | 1.8e-35 | 0.9893617021276596 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395098 | Cellulase | 2.88e-08 | 85 | 225 | 27 | 167 | Cellulase (glycosyl hydrolase family 5). |
| 367362 | Glutenin_hmw | 0.001 | 393 | 453 | 287 | 347 | High molecular weight glutenin subunit. Members of this family include high molecular weight subunits of glutenin. This group of gluten proteins is thought to be largely responsible for the elastic properties of gluten, and hence, doughs. Indeed, glutenin high molecular weight subunits are classified as elastomeric proteins, because the glutenin network can withstand significant deformations without breaking, and return to the original conformation when the stress is removed. Elastomeric proteins differ considerably in amino acid sequence, but they are all polymers whose subunits consist of elastomeric domains, composed of repeated motifs, and non-elastic domains that mediate cross-linking between the subunits. The elastomeric domain motifs are all rich in glycine residues in addition to other hydrophobic residues. High molecular weight glutenin subunits have an extensive central elastomeric domain, flanked by two terminal non-elastic domains that form disulphide cross-links. The central elastomeric domain is characterized by the following three repeated motifs: PGQGQQ, GYYPTS[P/L]QQ, GQQ. It possesses overlapping beta-turns within and between the repeated motifs, and assumes a regular helical secondary structure with a diameter of approx. 1.9 nm and a pitch of approx. 1.5 nm. |
| 225344 | BglC | 0.003 | 55 | 222 | 51 | 221 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
| 367362 | Glutenin_hmw | 0.006 | 369 | 453 | 509 | 589 | High molecular weight glutenin subunit. Members of this family include high molecular weight subunits of glutenin. This group of gluten proteins is thought to be largely responsible for the elastic properties of gluten, and hence, doughs. Indeed, glutenin high molecular weight subunits are classified as elastomeric proteins, because the glutenin network can withstand significant deformations without breaking, and return to the original conformation when the stress is removed. Elastomeric proteins differ considerably in amino acid sequence, but they are all polymers whose subunits consist of elastomeric domains, composed of repeated motifs, and non-elastic domains that mediate cross-linking between the subunits. The elastomeric domain motifs are all rich in glycine residues in addition to other hydrophobic residues. High molecular weight glutenin subunits have an extensive central elastomeric domain, flanked by two terminal non-elastic domains that form disulphide cross-links. The central elastomeric domain is characterized by the following three repeated motifs: PGQGQQ, GYYPTS[P/L]QQ, GQQ. It possesses overlapping beta-turns within and between the repeated motifs, and assumes a regular helical secondary structure with a diameter of approx. 1.9 nm and a pitch of approx. 1.5 nm. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.55e-26 | 47 | 357 | 23 | 325 | |
| 8.43e-26 | 47 | 359 | 23 | 327 | |
| 3.82e-25 | 47 | 351 | 95 | 396 | |
| 3.82e-25 | 47 | 351 | 95 | 396 | |
| 9.76e-25 | 44 | 368 | 93 | 408 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.85e-20 | 47 | 345 | 24 | 316 | Structrue of a lucidum protein [Ganoderma lucidum],5D8W_B Structrue of a lucidum protein [Ganoderma lucidum],5D8Z_A Structrue of a lucidum protein [Ganoderma lucidum],5D8Z_B Structrue of a lucidum protein [Ganoderma lucidum] |
|
| 6.41e-15 | 47 | 343 | 24 | 304 | Crystal structure of the endo-beta-1,4-glucanase (Xac0030) from Xanthomonas axonopodis pv. citri with the triple mutation His174Trp, Tyr211Ala and Lys227Arg. [Xanthomonas citri pv. citri str. 306],5HNN_B Crystal structure of the endo-beta-1,4-glucanase (Xac0030) from Xanthomonas axonopodis pv. citri with the triple mutation His174Trp, Tyr211Ala and Lys227Arg. [Xanthomonas citri pv. citri str. 306],5HNN_C Crystal structure of the endo-beta-1,4-glucanase (Xac0030) from Xanthomonas axonopodis pv. citri with the triple mutation His174Trp, Tyr211Ala and Lys227Arg. [Xanthomonas citri pv. citri str. 306] |
|
| 1.48e-14 | 47 | 343 | 24 | 304 | Crystal structure of XacCel5A in the native form [Xanthomonas citri pv. citri str. 306],4W7V_A Crystal structure of XacCel5A in complex with cellobiose [Xanthomonas citri pv. citri str. 306],4W7W_A High-resolution structure of XacCel5A in complex with cellopentaose [Xanthomonas citri pv. citri str. 306] |
|
| 1.55e-14 | 47 | 343 | 24 | 304 | Structure of XacCel5A crystallized in the space group P41212 [Xanthomonas citri pv. citri str. 306] |
|
| 3.23e-08 | 47 | 343 | 29 | 309 | Crystal structure of the endo-beta-1,4-glucanase Xac0029 from Xanthomonas axonopodis pv. citri [Xanthomonas citri pv. citri str. 306] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.57e-21 | 21 | 380 | 53 | 386 | Manganese dependent endoglucanase Eg5A OS=Phanerodontia chrysosporium OX=2822231 GN=Eg5A PE=1 SV=2 |
|
| 8.82e-20 | 44 | 359 | 32 | 329 | Endoglucanase 1 OS=Saitozyma flava OX=5416 GN=CMC1 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000228 | 0.999740 | CS pos: 23-24. Pr: 0.9764 |