You are browsing environment: FUNGIDB
CAZyme Information: PADG_03299-t30_1-p1
You are here: Home > Sequence: PADG_03299-t30_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
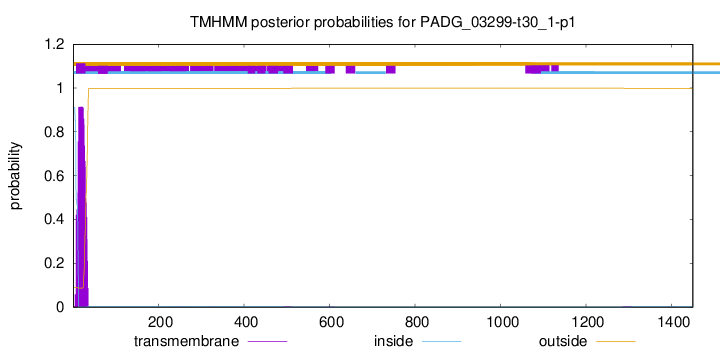
TMHMM annotations
Basic Information help
| Species | Paracoccidioides brasiliensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; NA; Paracoccidioides; Paracoccidioides brasiliensis | |||||||||||
| CAZyme ID | PADG_03299-t30_1-p1 | |||||||||||
| CAZy Family | GH16 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 3038048; End:3043395 Strand: - | |||||||||||
Full Sequence Download help
| MATRSHGLLQ ALLIWTLLSL QILASVMSPV LINPLRNAHE NKMHDLVPRN QLGTNVVLPP | 60 |
| SVAKATQLDI DEARRIVDDA IKEASALNKA RLEHPLRNHY QLKPGTGTSR RGEPNGDAPP | 120 |
| PPLFQISKEI AAAAALIAEV EAVAETRDSS KSKADYSYID VMYNRTVVRR ADAFWMEDIE | 180 |
| RKGAWPFGND RSFKVFRNVK DYGAVGNGVN DDTEAIKRAI ADTGTCGEKC NGATTKNSIV | 240 |
| YFPSGTYLVS STIETHFGSQ VIGNANDRPV IKAASSFIGL GVLSTNHYVE NGGNGTDGNA | 300 |
| KEWYVNTANF YRQIRNFRID ITATNQGAYI AALHYQVAQA TSLSNIDFIV SSNPGTTQQA | 360 |
| IFSENGSGGI LSDLTFTGGN FGIYGGNQQF TAQRLQFTNC KTAVQLIWDW GWIWKNIQIT | 420 |
| GSTTGFKLMS EDNVPRSGSI MVLDSIFRNT DTALLTFPAK QERAKGTTGI TLDNVAFDKV | 480 |
| RNAVADNQGK VYLAGSVGSI NTWALGPVYF DSSQRDFTLG MSFDTSREST LLADRLSALP | 540 |
| KAPFFERPRP QYEGVPVSDF VHMKDYAKGD GVTDDTEAFQ RVLREHSTDR IIFVDAGSYI | 600 |
| LTDTIIIPVG ARIVGEGWSQ LVASGPNFQD ERAPRSLVLV GQPGDTGEVE IQDLLFTTKG | 660 |
| PTAGAVLVEW NIKASSPGSA GMWDSHVRIG GASGTKLTST ECPAIRNGVN SDDCKSGSLM | 720 |
| MHITESSSAY MENVWLWVAD HDIDDPHLQD DNNTMVQTSV YSARGLLVES AEATWLYGTS | 780 |
| SEHAIYYQYN FYKAKNVFAG MIQTESPYYQ PTPKPPAPFE GAIGVLPGDP DFGGCRDGTP | 840 |
| GCDASWGLRI INSSQTYIAG AGLYSWFTTY TQECVDKRNC QNALIELKGN GPRVRIHNLI | 900 |
| TIGATNMLTS DGSEVPSQDN LAVDYHPYWS QVTVIDPFQN RGARGLTTPT SPDKSGRQCP | 960 |
| NIPPEVNVPG GKYPPDIPVM GRPGKSDHGY FTLVNGSPYN WMLTYNHSYQ LDQWKWHNVP | 1020 |
| AGESIQGEWK FARASNRYDD KGEAYYKIDG TDKSFQIWAR FYEDDPENAF HLRVLYDGLE | 1080 |
| TKDVKRGTSL DYRTRGGYGG MRAINWVLTG SEEEGYWSSH SPPVAWMSSI LNIIGDRKVK | 1140 |
| HVCMPGSHDA GMSKLDGHTQ FADEGNTLTQ YLSVYDQLRR GSRYFDVRPA IGNGGKYLTG | 1200 |
| HYSYVDVLGI GWQGGNGESI QEIVDGINRF TAENPELIII NLDLTLDTDN GYKPFNDEQW | 1260 |
| SKTFDLFEGV KYLQGNLEGD LTERTMNDYI GAGGAAVIVI ASGGPTRPEK GIYSNRQFPR | 1320 |
| YDSYSNSDDP AAMADDQLAK LKGNRNIVSD TTERKDTFHI FSWTLTLSRV FERTIADQSI | 1380 |
| ELGYDPLFWR GYHAFTPFSY PNVVYMDFIG SADQSETTLE KTHGEVTALA MAVNMELASR | 1440 |
| NCYVGGGTMV | 1450 |
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH55 | 168 | 920 | 2e-254 | 0.9716216216216216 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 176558 | PI-PLCXDc_like_2 | 1.69e-115 | 1131 | 1434 | 1 | 300 | Catalytic domain of uncharacterized hypothetical proteins similar to eukaryotic phosphatidylinositol-specific phospholipase C, X domain containing proteins. This subfamily corresponds to the catalytic domain present in a group of uncharacterized hypothetical proteins found in bacteria and fungi, which are similar to eukaryotic phosphatidylinositol-specific phospholipase C, X domain containing proteins (PI-PLCXD). The typical eukaryotic phosphoinositide-specific phospholipase C (PI-PLC, EC 3.1.4.11) has a multidomain organization that consists of a PLC catalytic core domain, and various regulatory domains. The catalytic core domain is assembled from two highly conserved X- and Y-regions split by a divergent linker sequence. In contrast, eukaryotic PI-PLCXDs contain a single TIM-barrel type catalytic domain, X domain, and are more closely related to bacterial PI-PLCs, which participate in Ca2+-independent PI metabolism, hydrolyzing the membrane lipid phosphatidylinositol (PI) to produce phosphorylated myo-inositol and diacylglycerol (DAG). Although the biological function of eukaryotic PI-PLCXDs still remains unclear, it may distinct from that of typical eukaryotic PI-PLCs. |
| 403800 | Pectate_lyase_3 | 2.01e-81 | 196 | 427 | 1 | 213 | Pectate lyase superfamily protein. This family of proteins possesses a beta helical structure like Pectate lyase. This family is most closely related to glycosyl hydrolase family 28. |
| 176529 | PI-PLCXDc_like | 2.43e-53 | 1132 | 1434 | 2 | 288 | Catalytic domain of phosphatidylinositol-specific phospholipase C X domain containing and similar proteins. This family corresponds to the catalytic domain present in phosphatidylinositol-specific phospholipase C X domain containing proteins (PI-PLCXD) which are bacterial phosphatidylinositol-specific phospholipase C (PI-PLC, EC 4.6.1.13) sequence homologs mainly found in eukaryota. The typical eukaryotic phosphoinositide-specific phospholipase C (PI-PLC, EC 3.1.4.11) have a multidomain organization that consists of a PLC catalytic core domain, and various regulatory domains. The catalytic core domain is assembled from two highly conserved X- and Y-regions split by a divergent linker sequence. In contrast, eukaryotic PI-PLCXDs and their bacterial homologs contain a single TIM-barrel type catalytic domain, X domain, which is more closely related to that of bacterial PI-PLCs. Although the biological function of eukaryotic PI-PLCXDs still remains unclear, it may be distinct from that of typical eukaryotic PI-PLCs. |
| 176500 | PI-PLCc_bacteria_like | 7.08e-32 | 1132 | 1413 | 2 | 262 | Catalytic domain of bacterial phosphatidylinositol-specific phospholipase C and similar proteins. This subfamily corresponds to the catalytic domain present in bacterial phosphatidylinositol-specific phospholipase C (PI-PLC, EC 4.6.1.13) and their sequence homologs found in eukaryota. Bacterial PI-PLCs participate in Ca2+-independent PI metabolism, hydrolyzing the membrane lipid phosphatidylinositol (PI) to produce phosphorylated myo-inositol and diacylglycerol (DAG). Although their precise physiological function remains unclear, bacterial PI-PLCs may function as virulence factors in some pathogenic bacteria. Bacterial PI-PLCs contain a single TIM-barrel type catalytic domain. Its catalytic mechanism is based on general base and acid catalysis utilizing two well conserved histidines, and consists of two steps, a phosphotransfer and a phosphodiesterase reaction. Eukaryotic homologs in this family are named as phosphatidylinositol-specific phospholipase C X domain containing proteins (PI-PLCXD). They are distinct from the typical eukaryotic phosphoinositide-specific phospholipases C (PI-PLC, EC 3.1.4.11), which have a multidomain organization that consists of a PLC catalytic core domain, and various regulatory domains. The catalytic core domain is assembled from two highly conserved X- and Y-regions split by a divergent linker sequence. In contrast, eukaryotic PI-PLCXDs contain a single TIM-barrel type catalytic domain, X domain, which is closely related to that of bacterial PI-PLCs. Although the biological function of eukaryotic PI-PLCXDs still remains unclear, it may be distinct from that of typical eukaryotic PI-PLCs. This family also includes a distinctly different type of eukaryotic PLC, glycosylphosphatidylinositol-specific phospholipase C (GPI-PLC), an integral membrane protein characterized in the protozoan parasite Trypanosoma brucei. T. brucei GPI-PLC hydrolyzes the GPI-anchor on the variant specific glycoprotein (VSG), releasing dimyristyl glycerol (DMG), which may facilitate the evasion of the protozoan to the host's immune system. It does not require Ca2+ for its activity and is more closely related to bacterial PI-PLCs, but not mammalian PI-PLCs. |
| 176557 | PI-PLCXDc_like_1 | 6.20e-11 | 1144 | 1335 | 14 | 185 | Catalytic domain of uncharacterized hypothetical proteins similar to eukaryotic phosphatidylinositol-specific phospholipase C, X domain containing proteins. This subfamily corresponds to the catalytic domain present in a group of uncharacterized hypothetical proteins found in bacteria and fungi, which are similar to eukaryotic phosphatidylinositol-specific phospholipase C, X domain containing proteins (PI-PLCXD). The typical eukaryotic phosphoinositide-specific phospholipase C (PI-PLC, EC 3.1.4.11) has a multidomain organization that consists of a PLC catalytic core domain, and various regulatory domains. The catalytic core domain is assembled from two highly conserved X- and Y-regions split by a divergent linker sequence. In contrast, eukaryotic PI-PLCXDs contain a single TIM-barrel type catalytic domain, X domain, and are more closely related to bacterial PI-PLCs, which participate in Ca2+-independent PI metabolism, hydrolyzing the membrane lipid phosphatidylinositol (PI) to produce phosphorylated myo-inositol and diacylglycerol (DAG). Although the biological function of eukaryotic PI-PLCXDs still remains unclear, it may distinct from that of typical eukaryotic PI-PLCs. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QSS51941.1|GH55 | 0.0 | 1 | 1450 | 1 | 1446 |
| QSS69923.1|GH55 | 0.0 | 345 | 1450 | 2 | 1107 |
| QBZ61798.1|GH55 | 0.0 | 41 | 1447 | 41 | 1428 |
| UKZ47543.1|GH55 | 0.0 | 62 | 938 | 64 | 932 |
| UKZ74106.1|GH55 | 0.0 | 62 | 938 | 64 | 932 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5M5Z_A | 7.90e-182 | 174 | 915 | 6 | 736 | Chain A, Beta-1,3-glucanase [Thermochaetoides thermophila],5M60_A Chain A, Beta-1,3-glucanase [Thermochaetoides thermophila] |
| 3EQN_A | 3.06e-169 | 172 | 920 | 24 | 743 | Chain A, Glucan 1,3-beta-glucosidase [Phanerodontia chrysosporium],3EQN_B Chain B, Glucan 1,3-beta-glucosidase [Phanerodontia chrysosporium],3EQO_A Chain A, Glucan 1,3-beta-glucosidase [Phanerodontia chrysosporium],3EQO_B Chain B, Glucan 1,3-beta-glucosidase [Phanerodontia chrysosporium] |
| 7CHU_A | 2.25e-06 | 141 | 273 | 27 | 149 | Chain A, Putative pectin lyase [Geobacillus virus E2],7CHU_B Chain B, Putative pectin lyase [Geobacillus virus E2],7CHU_C Chain C, Putative pectin lyase [Geobacillus virus E2] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| sp|D4B0V1|E13B_ARTBC | 1.90e-169 | 170 | 920 | 39 | 862 | Probable glucan endo-1,3-beta-glucosidase ARB_02077 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_02077 PE=1 SV=1 |
| sp|P49426|EXG1_COCCA | 8.04e-144 | 172 | 923 | 46 | 776 | Glucan 1,3-beta-glucosidase OS=Cochliobolus carbonum OX=5017 GN=EXG1 PE=1 SV=1 |
| sp|P53626|E13B_TRIHA | 3.68e-51 | 167 | 927 | 30 | 752 | Glucan endo-1,3-beta-glucosidase BGN13.1 OS=Trichoderma harzianum OX=5544 GN=bgn13.1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.058814 | 0.941165 | CS pos: 28-29. Pr: 0.3242 |