You are browsing environment: FUNGIDB
CAZyme Information: PABG_01806-t30_1-p1
You are here: Home > Sequence: PABG_01806-t30_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
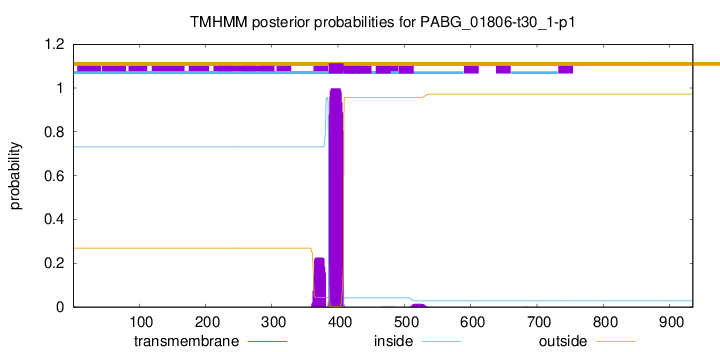
TMHMM annotations
Basic Information help
| Species | Paracoccidioides brasiliensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; NA; Paracoccidioides; Paracoccidioides brasiliensis | |||||||||||
| CAZyme ID | PABG_01806-t30_1-p1 | |||||||||||
| CAZy Family | GH125 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.58:2 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 559 | 893 | 6.3e-105 | 0.9966996699669967 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 225344 | BglC | 1.14e-44 | 510 | 911 | 18 | 383 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
| 395098 | Cellulase | 8.87e-09 | 564 | 794 | 24 | 236 | Cellulase (glycosyl hydrolase family 5). |
| 237171 | PRK12678 | 5.54e-05 | 69 | 240 | 56 | 220 | transcription termination factor Rho; Provisional |
| 237171 | PRK12678 | 0.003 | 87 | 244 | 125 | 276 | transcription termination factor Rho; Provisional |
| 273727 | U2AF_lg | 0.003 | 175 | 255 | 1 | 70 | U2 snRNP auxilliary factor, large subunit, splicing factor. These splicing factors consist of an N-terminal arginine-rich low complexity domain followed by three tandem RNA recognition motifs (pfam00076). The well-characterized members of this family are auxilliary components of the U2 small nuclear ribonuclearprotein splicing factor (U2AF). These proteins are closely related to the CC1-like subfamily of splicing factors (TIGR01622). Members of this subfamily are found in plants, metazoa and fungi. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 935 | 1 | 948 | |
| 5.15e-305 | 465 | 935 | 1 | 471 | |
| 7.30e-305 | 465 | 935 | 1 | 471 | |
| 1.98e-216 | 437 | 935 | 347 | 830 | |
| 4.91e-216 | 437 | 935 | 65 | 549 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.21e-64 | 500 | 918 | 5 | 393 | Exo-B-(1,3)-Glucanase from Candida Albicans in complex with unhydrolysed and covalently linked 2,4-dinitrophenyl-2-deoxy-2-fluoro-B-D-glucopyranoside at 1.9 A [Candida albicans] |
|
| 9.61e-64 | 503 | 918 | 2 | 387 | Exo-b-(1,3)-glucanase From Candida Albicans [Candida albicans] |
|
| 1.52e-63 | 500 | 918 | 4 | 392 | F144Y/F258Y Double Mutant of Exo-beta-1,3-glucanase from Candida albicans at 2 A [Candida albicans] |
|
| 1.82e-63 | 503 | 918 | 2 | 387 | Exo-b-(1,3)-glucanase From Candida Albicans At 1.85 A Resolution [Candida albicans],1EQC_A Exo-b-(1,3)-glucanase From Candida Albicans In Complex With Castanospermine At 1.85 A [Candida albicans] |
|
| 2.14e-63 | 500 | 918 | 5 | 393 | Chain A, Hypothetical protein XOG1 [Candida albicans] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.35e-238 | 249 | 935 | 175 | 834 | Probable glucan 1,3-beta-glucosidase D OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=exgD PE=3 SV=1 |
|
| 1.14e-236 | 253 | 935 | 178 | 833 | Probable glucan 1,3-beta-glucosidase D OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=exgD PE=3 SV=1 |
|
| 1.14e-236 | 253 | 935 | 178 | 833 | Probable glucan 1,3-beta-glucosidase D OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=exgD PE=3 SV=1 |
|
| 1.48e-233 | 249 | 935 | 172 | 830 | Probable glucan 1,3-beta-glucosidase D OS=Aspergillus clavatus (strain ATCC 1007 / CBS 513.65 / DSM 816 / NCTC 3887 / NRRL 1 / QM 1276 / 107) OX=344612 GN=exgD PE=3 SV=1 |
|
| 3.51e-217 | 437 | 935 | 347 | 830 | Probable glucan 1,3-beta-glucosidase D OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=exgD PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000069 | 0.000000 |