You are browsing environment: FUNGIDB
CAZyme Information: PAAG_05717-t30_1-p1
You are here: Home > Sequence: PAAG_05717-t30_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Paracoccidioides lutzii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; NA; Paracoccidioides; Paracoccidioides lutzii | |||||||||||
| CAZyme ID | PAAG_05717-t30_1-p1 | |||||||||||
| CAZy Family | GH76 | |||||||||||
| CAZyme Description | glycosyl hydrolase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH76 | 32 | 378 | 4.6e-89 | 0.9022346368715084 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 397638 | Glyco_hydro_76 | 5.93e-30 | 124 | 357 | 83 | 331 | Glycosyl hydrolase family 76. Family of alpha-1,6-mannanases. |
| 227170 | COG4833 | 6.20e-16 | 123 | 371 | 88 | 338 | Predicted alpha-1,6-mannanase, GH76 family [Carbohydrate transport and metabolism]. |
| 395616 | Glyco_hydro_9 | 0.001 | 120 | 212 | 203 | 284 | Glycosyl hydrolase family 9. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 6.59e-187 | 13 | 383 | 16 | 386 | |
| 2.19e-185 | 20 | 383 | 23 | 386 | |
| 7.13e-166 | 20 | 383 | 26 | 370 | |
| 8.31e-133 | 21 | 383 | 31 | 380 | |
| 8.31e-133 | 21 | 383 | 31 | 380 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.75e-19 | 124 | 358 | 107 | 340 | Chain A, Alpha-1,6-mannanase [Niallia circulans],4D4A_B Chain B, Alpha-1,6-mannanase [Niallia circulans],4D4B_A Chain A, Alpha-1,6-mannanase [Niallia circulans],4D4B_B Chain B, Alpha-1,6-mannanase [Niallia circulans],4D4C_A Chain A, Alpha-1,6-mannanase [Niallia circulans],4D4C_B Chain B, Alpha-1,6-mannanase [Niallia circulans],4D4D_A Chain A, Alpha-1,6-mannanase [Niallia circulans],4D4D_B Chain B, Alpha-1,6-mannanase [Niallia circulans],5N0F_A Chain A, Alpha-1,6-mannanase [Niallia circulans],5N0F_B Chain B, Alpha-1,6-mannanase [Niallia circulans],6ZBX_A Chain A, Alpha-1,6-mannanase [Niallia circulans],6ZBX_B Chain B, Alpha-1,6-mannanase [Niallia circulans],7NL5_A Chain A, Alpha-1,6-mannanase [Niallia circulans] |
|
| 3.52e-19 | 124 | 358 | 86 | 319 | Chain A, Alpha-1,6-mannanase [Niallia circulans] |
|
| 3.64e-19 | 124 | 358 | 89 | 322 | Chain A, Alpha-1,6-mannanase [Niallia circulans],4BOJ_B Chain B, Alpha-1,6-mannanase [Niallia circulans],4BOJ_C Chain C, Alpha-1,6-mannanase [Niallia circulans] |
|
| 5.05e-19 | 124 | 358 | 120 | 353 | Chain A, Alpha-1,6-mannanase [Niallia circulans],5M77_B Chain B, Alpha-1,6-mannanase [Niallia circulans] |
|
| 8.07e-19 | 124 | 358 | 107 | 340 | Chain A, Alpha-1,6-mannanase [Niallia circulans],5AGD_B Chain B, Alpha-1,6-mannanase [Niallia circulans],6ZBM_A Chain A, Alpha-1,6-mannanase [Niallia circulans],6ZBW_A Chain A, Alpha-1,6-mannanase [Niallia circulans],6ZBW_B Chain B, Alpha-1,6-mannanase [Niallia circulans] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.71e-18 | 48 | 357 | 65 | 365 | Mannan endo-1,6-alpha-mannosidase DFG5 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=DFG5 PE=1 SV=1 |
|
| 2.84e-16 | 128 | 317 | 110 | 310 | Putative mannan endo-1,6-alpha-mannosidase C1198.06c OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=SPBC1198.06c PE=3 SV=1 |
|
| 2.31e-12 | 128 | 356 | 122 | 367 | Mannan endo-1,6-alpha-mannosidase DFG5 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=DFG5 PE=1 SV=1 |
|
| 9.58e-12 | 128 | 356 | 117 | 360 | Mannan endo-1,6-alpha-mannosidase DCW1 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=DCW1 PE=1 SV=1 |
|
| 1.69e-11 | 128 | 356 | 114 | 357 | Mannan endo-1,6-alpha-mannosidase DCW1 OS=Candida glabrata (strain ATCC 2001 / CBS 138 / JCM 3761 / NBRC 0622 / NRRL Y-65) OX=284593 GN=DCW1 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
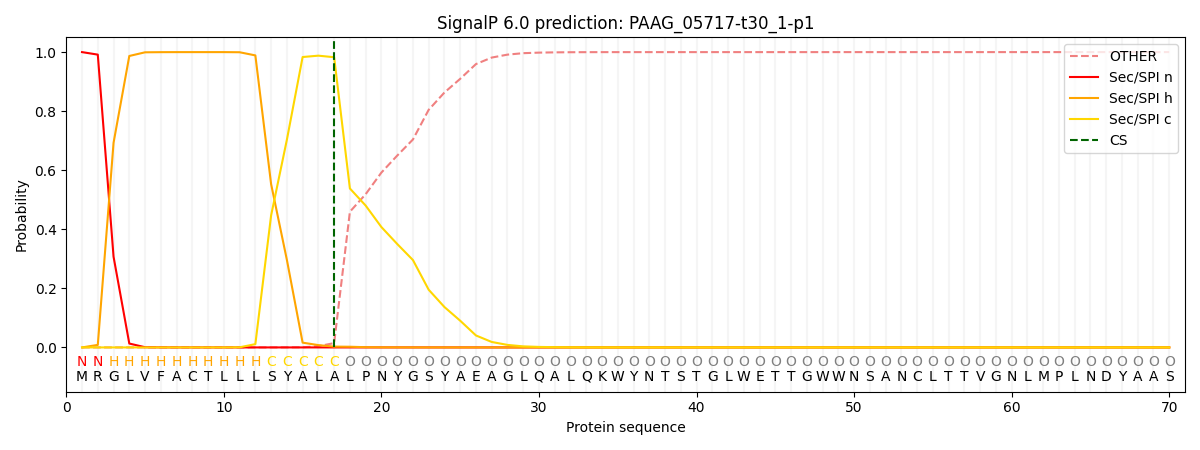
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000239 | 0.999730 | CS pos: 17-18. Pr: 0.9826 |
