You are browsing environment: FUNGIDB
CAZyme Information: P170DRAFT_419063-t37_1-p1
You are here: Home > Sequence: P170DRAFT_419063-t37_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus steynii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus steynii | |||||||||||
| CAZyme ID | P170DRAFT_419063-t37_1-p1 | |||||||||||
| CAZy Family | GH2 | |||||||||||
| CAZyme Description | putative beta-glucosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.120:1 | 3.2.1.21:1 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 113 | 337 | 9.1e-62 | 0.9768518518518519 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 185053 | PRK15098 | 3.55e-172 | 31 | 775 | 26 | 762 | beta-glucosidase BglX. |
| 224389 | BglX | 9.55e-80 | 47 | 478 | 1 | 396 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| 178629 | PLN03080 | 4.73e-73 | 116 | 737 | 79 | 742 | Probable beta-xylosidase; Provisional |
| 396478 | Glyco_hydro_3_C | 2.80e-61 | 411 | 660 | 1 | 216 | Glycosyl hydrolase family 3 C-terminal domain. This domain is involved in catalysis and may be involved in binding beta-glucan. This domain is found associated with pfam00933. |
| 395747 | Glyco_hydro_3 | 1.01e-59 | 48 | 371 | 1 | 316 | Glycosyl hydrolase family 3 N terminal domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 778 | 1 | 779 | |
| 0.0 | 1 | 778 | 1 | 779 | |
| 0.0 | 1 | 778 | 1 | 779 | |
| 0.0 | 1 | 778 | 1 | 779 | |
| 0.0 | 1 | 778 | 1 | 779 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 0.0 | 26 | 778 | 4 | 757 | Isoprimeverose-producing enzyme from Aspergillus oryzae in complex with isoprimeverose [Aspergillus oryzae RIB40],5YOT_B Isoprimeverose-producing enzyme from Aspergillus oryzae in complex with isoprimeverose [Aspergillus oryzae RIB40],5YQS_A Isoprimeverose-producing enzyme from Aspergillus oryzae in complex with isoprimeverose [Aspergillus oryzae RIB40],5YQS_B Isoprimeverose-producing enzyme from Aspergillus oryzae in complex with isoprimeverose [Aspergillus oryzae RIB40] |
|
| 0.0 | 26 | 778 | 4 | 757 | Chain A, Fn3_like domain-containing protein [Aspergillus oryzae RIB40] |
|
| 2.66e-121 | 40 | 770 | 13 | 744 | Crystal Structure of Glycosil Hydrolase Family 3 N-Terminal Domain Protein from Bacteroides intestinalis [Bacteroides intestinalis DSM 17393],5TF0_B Crystal Structure of Glycosil Hydrolase Family 3 N-Terminal Domain Protein from Bacteroides intestinalis [Bacteroides intestinalis DSM 17393] |
|
| 9.18e-115 | 100 | 770 | 105 | 778 | Chain A, EmGH1 [Aurantiacibacter marinus],5Z87_B Chain B, EmGH1 [Aurantiacibacter marinus] |
|
| 1.03e-113 | 40 | 770 | 14 | 745 | Crystal structure of GH3 beta-glucosidase from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482],5XXL_B Crystal structure of GH3 beta-glucosidase from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482],5XXM_A Crystal structure of GH3 beta-glucosidase from Bacteroides thetaiotaomicron in complex with gluconolactone [Bacteroides thetaiotaomicron VPI-5482],5XXM_B Crystal structure of GH3 beta-glucosidase from Bacteroides thetaiotaomicron in complex with gluconolactone [Bacteroides thetaiotaomicron VPI-5482] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.12e-124 | 15 | 770 | 4 | 758 | Periplasmic beta-glucosidase OS=Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) OX=99287 GN=bglX PE=3 SV=2 |
|
| 4.74e-122 | 40 | 770 | 38 | 758 | Periplasmic beta-glucosidase OS=Escherichia coli (strain K12) OX=83333 GN=bglX PE=3 SV=2 |
|
| 5.35e-119 | 113 | 767 | 60 | 711 | Beta-xylosidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22130 PE=1 SV=1 |
|
| 4.93e-106 | 29 | 770 | 29 | 748 | Putative beta-xylosidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22230 PE=2 SV=1 |
|
| 1.69e-99 | 116 | 778 | 35 | 675 | Thermostable beta-glucosidase B OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=bglB PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
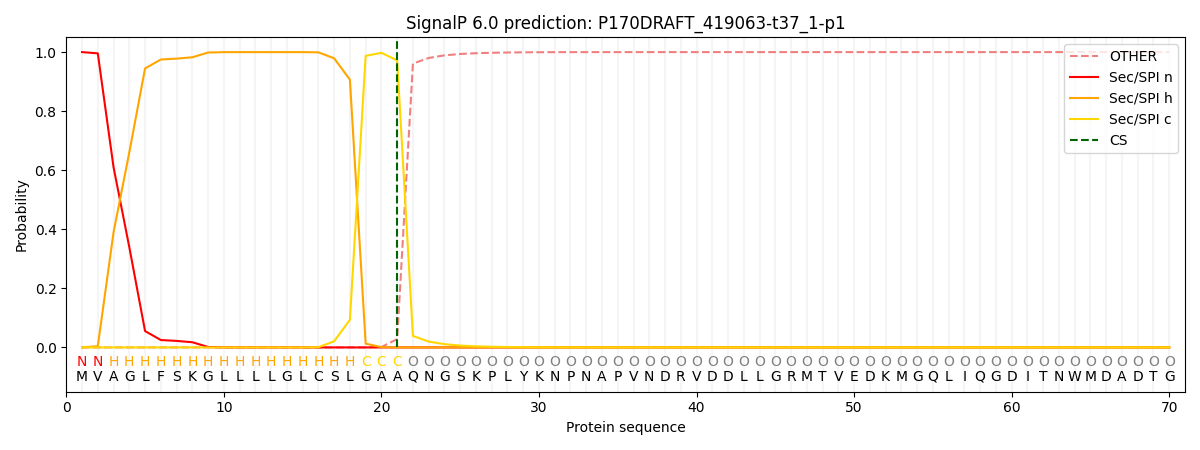
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000196 | 0.999785 | CS pos: 21-22. Pr: 0.9720 |
