You are browsing environment: FUNGIDB
CAZyme Information: OTA36282.1
You are here: Home > Sequence: OTA36282.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Hortaea werneckii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Dothideomycetes; ; Teratosphaeriaceae; Hortaea; Hortaea werneckii | |||||||||||
| CAZyme ID | OTA36282.1 | |||||||||||
| CAZy Family | GH79 | |||||||||||
| CAZyme Description | Beta-galactosidase [Source:UniProtKB/TrEMBL;Acc:A0A1Z5TJQ8] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH35 | 40 | 387 | 1.6e-81 | 0.9869706840390879 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396048 | Glyco_hydro_35 | 1.94e-90 | 39 | 387 | 1 | 314 | Glycosyl hydrolases family 35. |
| 402180 | BetaGal_dom2 | 3.71e-61 | 397 | 577 | 1 | 180 | Beta-galactosidase, domain 2. This is the second domain of the five-domain beta-galactosidase enzyme that altogether catalyzes the hydrolysis of beta(1-3) and beta(1-4) galactosyl bonds in oligosaccharides as well as the inverse reaction of enzymatic condensation and trans-glycosylation. This domain is made up of 16 antiparallel beta-strands and an alpha-helix at its C-terminus. The fold of this domain appears to be unique. In addition, the last seven strands of the domain form a subdomain with an immunoglobulin-like (I-type Ig) fold in which the first strand is divided between the two beta-sheets. In penicillin spp this strand is interrupted by a 12-residue insertion which forms an additional edge-strand to the second beta-sheet of the sub-domain. The remainder of the second domain forms a series of beta-hairpins at its N-terminus, four strands of which are contiguous with part of the Ig-like sub-domain, forming in total a seven-stranded antiparallel beta-sheet. This domain is associated with family Glyco_hydro_35, pfam01301, which is N-terminal to it, but itself has no metazoan members. |
| 198097 | BetaGal_dom2 | 1.90e-51 | 396 | 577 | 1 | 181 | Beta-galactosidase, domain 2. This is the second domain of the five-domain beta-galactosidase enzyme that altogether catalyses the hydrolysis of beta(1-3) and beta(1-4) galactosyl bonds in oligosaccharides as well as the inverse reaction of enzymatic condensation and trans-glycosylation. This domain is made up of 16 antiparallel beta-strands and an alpha-helix at its C terminus. The fold of this domain appears to be unique. In addition, the last seven strands of the domain form a subdomain with an immunoglobulin-like (I-type Ig) fold in which the first strand is divided between the two beta-sheets. In penicillin spp this strand is interrupted by a 12-residue insertion which forms an additional edge-strand to the second beta-sheet of the sub-domain. The remainder of the second domain forms a series of beta-hairpins at its N terminus, four strands of which are contiguous with part of the Ig-like sub-domain, forming in total a seven-stranded antiparallel beta-sheet. This domain is associated with family Glyco_hydro_35, which is N-terminal to it, but itself has no metazoan members. |
| 404274 | BetaGal_dom4_5 | 3.73e-32 | 871 | 981 | 1 | 111 | Beta-galactosidase jelly roll domain. This domain is found in beta galactosidase enzymes. It has a jelly roll fold. |
| 166698 | PLN03059 | 4.04e-26 | 33 | 373 | 30 | 323 | beta-galactosidase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 14 | 1020 | 11 | 996 | |
| 0.0 | 14 | 1020 | 19 | 1002 | |
| 0.0 | 1 | 1020 | 1 | 1030 | |
| 0.0 | 1 | 1019 | 1 | 1004 | |
| 0.0 | 3 | 1019 | 1 | 1013 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.64e-182 | 4 | 985 | 19 | 974 | Structure Of Beta-galactosidase From Aspergillus Niger [Aspergillus niger CBS 513.88] |
|
| 2.68e-181 | 4 | 985 | 19 | 974 | STRUCTURE OF E298Q-BETA-GALACTOSIDASE FROM ASPERGILLUS NIGER IN COMPLEX WITH 3-b-Galactopyranosyl glucose [Aspergillus niger CBS 513.88],5IHR_A Structure Of E298q-beta-galactosidase From Aspergillus Niger In Complex With Allolactose [Aspergillus niger CBS 513.88],5JUV_A STRUCTURE OF E298Q-BETA-GALACTOSIDASE FROM ASPERGILLUS NIGER IN COMPLEX WITH 6-b-Galactopyranosyl galactose [Aspergillus niger CBS 513.88],5MGC_A STRUCTURE OF E298Q-BETA-GALACTOSIDASE FROM ASPERGILLUS NIGER IN COMPLEX WITH 4-Galactosyl-lactose [Aspergillus niger CBS 513.88],5MGD_A STRUCTURE OF E298Q-BETA-GALACTOSIDASE FROM ASPERGILLUS NIGER IN COMPLEX WITH 6-Galactosyl-lactose [Aspergillus niger CBS 513.88] |
|
| 4.57e-181 | 29 | 985 | 2 | 938 | Native structure of beta-galactosidase from Penicillium sp. [Penicillium sp.],1XC6_A Native Structure Of Beta-Galactosidase from Penicillium sp. in complex with Galactose [Penicillium sp.] |
|
| 5.28e-174 | 23 | 1001 | 36 | 993 | Crystal structure of beta-galactosidase from Aspergillus oryzae in complex with galactose [Aspergillus oryzae] |
|
| 1.27e-165 | 29 | 985 | 22 | 970 | Chain A, Beta-galactosidase [Trichoderma reesei],3OGR_A Chain A, Beta-galactosidase [Trichoderma reesei],3OGS_A Chain A, Beta-galactosidase [Trichoderma reesei],3OGV_A Chain A, Beta-galactosidase [Trichoderma reesei] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 0.0 | 2 | 1019 | 6 | 1016 | Probable beta-galactosidase B OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=lacB PE=3 SV=2 |
|
| 0.0 | 12 | 1019 | 19 | 1021 | Probable beta-galactosidase B OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=lacB PE=3 SV=2 |
|
| 0.0 | 21 | 1000 | 28 | 1003 | Probable beta-galactosidase B OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=lacB PE=3 SV=2 |
|
| 0.0 | 10 | 1019 | 12 | 1014 | Probable beta-galactosidase B OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=lacB PE=3 SV=1 |
|
| 0.0 | 22 | 1019 | 26 | 1007 | Probable beta-galactosidase B OS=Sclerotinia sclerotiorum (strain ATCC 18683 / 1980 / Ss-1) OX=665079 GN=lacB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
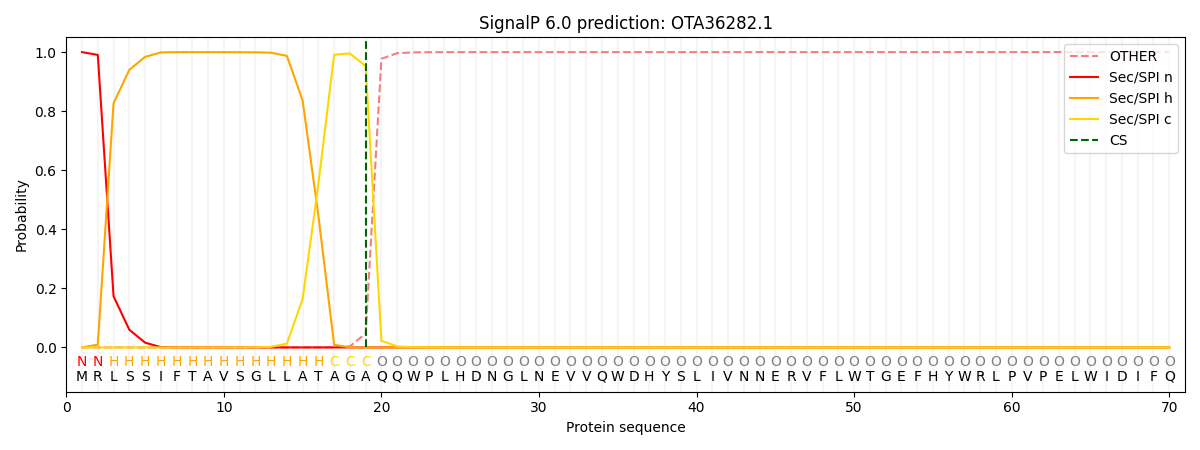
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000572 | 0.999397 | CS pos: 19-20. Pr: 0.9526 |
