You are browsing environment: FUNGIDB
CAZyme Information: OTA35730.1
You are here: Home > Sequence: OTA35730.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Hortaea werneckii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Dothideomycetes; ; Teratosphaeriaceae; Hortaea; Hortaea werneckii | |||||||||||
| CAZyme ID | OTA35730.1 | |||||||||||
| CAZy Family | GH72 | |||||||||||
| CAZyme Description | Non-reducing end alpha-L-arabinofuranosidase [Source:UniProtKB/TrEMBL;Acc:A0A1Z5TIA3] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 146792; End:153759 Strand: - | |||||||||||
Full Sequence Download help
| MHLKPVALAA ALAPFAAALD VQVASSGGNK TSYHQYGFLH EDINNSGDGG IYAELIVNRA | 60 |
| FQSSEMYPVS LNGYHSLHGA RLSIQNLSQP LSAALDSSMR VAAGNGSSRA GFENEGYWGM | 120 |
| NVECQKYTGS FWVRGAYDGS FTASLQSNLT DDVFGSVEVP SQAVADDWVE HTFTLTPDRD | 180 |
| APNSNNTFAI TFNPAGTENG YLDFNLISLF PPTYKDRPNG LRVDLAQVLE AYHPTVFRFP | 240 |
| GGNMLEGNTN ETYWDWKKTL GPLRERPGFP GVWGYMQTNG LGLMEYLEMA EDINNMTIGK | 300 |
| QRRLSRRQLS NVDHSLVVGV WAGLSLNGDI TPRNDLQPFI DDALNEIEFI RGPVDSPWGS | 360 |
| RRAELGHPEP FNLQYVEVGN EDWLAGAPEG WNSYKEYRFP MFMQAINDAY PDIQVIASGA | 420 |
| TSDNYTFPAP AIGDYHPYRE PDALVEEFNR FDNDLGHIVG EVSATFPNNG SKFDGPLFEF | 480 |
| PAWIGAVGSA ISMIGYERNA DRIPGTFYAP ILRNMNRWQW AVTMIQHAAD PALTTRSVSW | 540 |
| YIWEALAASP ISHTLPANEQ FGPVFYVAGR DENRHDALVW KGACYNISSN DIYNSTYRAN | 600 |
| STRVPVSVHF DGVEAGTEAT LTMVTNPGGD PYAMVDPLEG NEVVSTTETP ITANADGVFE | 660 |
| FYMTELSLAV LDTGVNKRVA GAPYSPISKS KHDWKPRMAG KPPGYRPSRY GEGAREFWQS | 720 |
| TLRGVKRSDS TVIDPIKHFL STELRAFKES RNALQATQKS FDHVIARYLS QSKTKEASSL | 780 |
| REDAFQVHEA RRAYIKASMD FCTGAPQVRA SLDKLIVRIF SDQWKDMRAS REANNGIFAK | 840 |
| LSSEMERVRG WSSEMDNSER AFKRELLTAR RQIEDSVSSR IRPSRELEDY SASTVPYLGS | 900 |
| GAGGATGITS KSANERLEQQ GWLFMKTTTG KPARTVWSRR WVFVKSGIFG WLIQGAKSGG | 960 |
| VEESEKIGVL LCSIRPAFQE DRRFCFEIKT KDTSIILQAE TQDELTEWIS ALEVAKKKAL | 1020 |
| EDPTSTETSP LSSGVDPAFA VSSPVAPEFA AKLGDSQPDE ARESEASVDQ RRITIKVFLE | 1080 |
| PLRLLQRRLS YLVRNANSED PDSLEEVLKV LIKLEADVPQ RSSSVESWED ITYEGGSPTT | 1140 |
| AGDQQKSPRR YGKNVKASLR IDGNLYGDPA RTGREVQKFK LPAQPVVYAP QGMQASVTRD | 1200 |
| FNVSAKALFH VMFGDKSAVF QLLYANRWAD KISQSPWSKP AEAKGPWTRK FTSQDQDSPM | 1260 |
| SDTQTVDVSN DHLCYVVTNN KNPWRLPYAT RFGLVTKIVI THTSKSKCKL AIFQQVNWLQ | 1320 |
| APQWSYIKHL IEKQALNSLE ADALDLTNVA MDQVAKLGNH SKTNRAIEIF GGIGQQLQVP | 1380 |
| QIDVSAVPNV ARSIAANGAR PQETTGLPRL VLDDVFAKTL QLVSMVIDIL IAVGKGAVGV | 1440 |
| CTAHTFLVAL LATSMIYNSW YGYRDSLLWY QERSAGKFMS RVGVTPDPVM GRSIYLRDLE | 1500 |
| ELVMPADVNE TLDAMAPSSS MAEAKTCRTT FSDMIAADDA SIATPQKHAA RLHNTRDAIA | 1560 |
| RYRHDLLIAL RVVNRVENDV LKAEWEDWVR SEERKCAKVE RMLQRGESGK KSSKSKLDVD | 1620 |
| AQLGEEFAEY CRSCRTEVAG LLNETRIM | 1648 |
Enzyme Prediction help
| EC | 3.2.1.55:10 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH51 | 206 | 542 | 2.5e-57 | 0.49047619047619045 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 153293 | BAR_SIP3_fungi | 2.39e-56 | 708 | 855 | 66 | 214 | The Bin/Amphiphysin/Rvs (BAR) domain of fungal Snf1p-interacting protein 3. BAR domains are dimerization, lipid binding and curvature sensing modules found in many different proteins with diverse functions including organelle biogenesis, membrane trafficking or remodeling, and cell division and migration. This group is composed of mostly uncharacterized fungal proteins with similarity to Saccharomyces cerevisiae Snf1p-interacting protein 3 (SIP3). These proteins contain an N-terminal BAR domain followed by a Pleckstrin Homology (PH) domain. SIP3 interacts with SNF1 protein kinase and activates transcription when anchored to DNA. It may function in the SNF1 pathway. BAR domains form dimers that bind to membranes, induce membrane bending and curvature, and may also be involved in protein-protein interactions. |
| 270098 | PH_SIP3 | 8.69e-55 | 918 | 1020 | 1 | 104 | Snf1p-interacting protein 3 Pleckstrin homology (PH) domain. SIP3 interacts with SNF1 protein kinase and activates transcription when anchored to DNA. It may function in the SNF1 pathway. SIP3 contain an N-terminal Bin/Amphiphysin/Rvs (BAR) domain followed by a PH domain. BAR domains form dimers that bind to membranes, induce membrane bending and curvature, and may also be involved in protein-protein interactions. PH domains have diverse functions, but in general are involved in targeting proteins to the appropriate cellular location or in the interaction with a binding partner. They share little sequence conservation, but all have a common fold, which is electrostatically polarized. Less than 10% of PH domains bind phosphoinositide phosphates (PIPs) with high affinity and specificity. PH domains are distinguished from other PIP-binding domains by their specific high-affinity binding to PIPs with two vicinal phosphate groups: PtdIns(3,4)P2, PtdIns(4,5)P2 or PtdIns(3,4,5)P3 which results in targeting some PH domain proteins to the plasma membrane. A few display strong specificity in lipid binding. Any specificity is usually determined by loop regions or insertions in the N-terminus of the domain, which are not conserved across all PH domains. PH domains are found in cellular signaling proteins such as serine/threonine kinase, tyrosine kinases, regulators of G-proteins, endocytotic GTPases, adaptors, as well as cytoskeletal associated molecules and in lipid associated enzymes. |
| 406424 | DUF4782 | 1.12e-26 | 1197 | 1336 | 3 | 145 | Domain of unknown function (DUF4782). This presumed domain is functionally uncharacterized. This domain family is found in eukaryotes, and is approximately 150 amino acids in length. The family is found in association with pfam02893. |
| 270070 | PH_ACAP | 4.81e-23 | 919 | 1021 | 1 | 98 | ArfGAP with coiled-coil, ankyrin repeat and PH domains Pleckstrin homology (PH) domain. ACAP (also called centaurin beta) functions both as a Rab35 effector and as an Arf6-GTPase-activating protein (GAP) by which it controls actin remodeling and membrane trafficking. ACAP contain an NH2-terminal bin/amphiphysin/Rvs (BAR) domain, a phospholipid-binding domain, a PH domain, a GAP domain, and four ankyrin repeats. The AZAPs constitute a family of Arf GAPs that are characterized by an NH2-terminal pleckstrin homology (PH) domain and a central Arf GAP domain followed by two or more ankyrin repeats. On the basis of sequence and domain organization, the AZAP family is further subdivided into four subfamilies: 1) the ACAPs contain an NH2-terminal bin/amphiphysin/Rvs (BAR) domain (a phospholipid-binding domain that is thought to sense membrane curvature), a single PH domain followed by the GAP domain, and four ankyrin repeats; 2) the ASAPs also contain an NH2-terminal BAR domain, the tandem PH domain/GAP domain, three ankyrin repeats, two proline-rich regions, and a COOH-terminal Src homology 3 domain; 3) the AGAPs contain an NH2-terminal GTPase-like domain (GLD), a split PH domain, and the GAP domain followed by four ankyrin repeats; and 4) the ARAPs contain both an Arf GAP domain and a Rho GAP domain, as well as an NH2-terminal sterile-a motif (SAM), a proline-rich region, a GTPase-binding domain, and five PH domains. PMID 18003747 and 19055940 Centaurin can bind to phosphatidlyinositol (3,4,5)P3. PH domains have diverse functions, but in general are involved in targeting proteins to the appropriate cellular location or in the interaction with a binding partner. They share little sequence conservation, but all have a common fold, which is electrostatically polarized. Less than 10% of PH domains bind phosphoinositide phosphates (PIPs) with high affinity and specificity. PH domains are distinguished from other PIP-binding domains by their specific high-affinity binding to PIPs with two vicinal phosphate groups: PtdIns(3,4)P2, PtdIns(4,5)P2 or PtdIns(3,4,5)P3 which results in targeting some PH domain proteins to the plasma membrane. A few display strong specificity in lipid binding. Any specificity is usually determined by loop regions or insertions in the N-terminus of the domain, which are not conserved across all PH domains. PH domains are found in cellular signaling proteins such as serine/threonine kinase, tyrosine kinases, regulators of G-proteins, endocytotic GTPases, adaptors, as well as cytoskeletal associated molecules and in lipid associated enzymes. |
| 226064 | AbfA | 2.60e-20 | 208 | 429 | 37 | 223 | Alpha-L-arabinofuranosidase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| UQC86406.1|GH51 | 4.42e-313 | 1 | 677 | 2 | 650 |
| QDO67168.1|GH51 | 1.89e-309 | 4 | 675 | 5 | 647 |
| UPK90404.1|GH51 | 1.59e-301 | 1 | 684 | 2 | 662 |
| UJO24967.1|GH51 | 5.82e-299 | 3 | 676 | 4 | 673 |
| QRD05473.1|GH51 | 8.37e-297 | 4 | 681 | 5 | 663 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6ZPS_AAA | 4.26e-94 | 36 | 558 | 18 | 531 | Chain AAA, MgGH51 [Meripilus giganteus],6ZPV_AAA Chain AAA, MgGH51 [Meripilus giganteus],6ZPW_AAA Chain AAA, MgGH51 [Meripilus giganteus],6ZPX_AAA Chain AAA, MgGH51 [Meripilus giganteus],6ZPY_AAA Chain AAA, MgGH51 [Meripilus giganteus],6ZPZ_AAA Chain AAA, MgGH51 [Meripilus giganteus],6ZQ0_AAA Chain AAA, MgGH51 [Meripilus giganteus],6ZQ1_AAA Chain AAA, MgGH51 [Meripilus giganteus] |
| 4ATW_A | 5.73e-09 | 220 | 419 | 48 | 211 | The crystal structure of Arabinofuranosidase [Thermotoga maritima MSB8],4ATW_B The crystal structure of Arabinofuranosidase [Thermotoga maritima MSB8],4ATW_C The crystal structure of Arabinofuranosidase [Thermotoga maritima MSB8],4ATW_D The crystal structure of Arabinofuranosidase [Thermotoga maritima MSB8],4ATW_E The crystal structure of Arabinofuranosidase [Thermotoga maritima MSB8],4ATW_F The crystal structure of Arabinofuranosidase [Thermotoga maritima MSB8] |
| 3S2C_A | 5.76e-09 | 220 | 419 | 48 | 211 | Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_B Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_C Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_D Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_E Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_F Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_G Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_H Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_I Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_J Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_K Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_L Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1] |
| 3UG3_A | 6.00e-09 | 220 | 419 | 68 | 231 | Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima ligand free form [Thermotoga maritima],3UG3_B Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima ligand free form [Thermotoga maritima],3UG3_C Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima ligand free form [Thermotoga maritima],3UG3_D Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima ligand free form [Thermotoga maritima],3UG3_E Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima ligand free form [Thermotoga maritima],3UG3_F Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima ligand free form [Thermotoga maritima],3UG4_A Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima arabinose complex [Thermotoga maritima],3UG4_B Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima arabinose complex [Thermotoga maritima],3UG4_C Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima arabinose complex [Thermotoga maritima],3UG4_D Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima arabinose complex [Thermotoga maritima],3UG4_E Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima arabinose complex [Thermotoga maritima],3UG4_F Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima arabinose complex [Thermotoga maritima],3UG5_A Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima xylose complex [Thermotoga maritima],3UG5_B Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima xylose complex [Thermotoga maritima],3UG5_C Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima xylose complex [Thermotoga maritima],3UG5_D Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima xylose complex [Thermotoga maritima],3UG5_E Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima xylose complex [Thermotoga maritima],3UG5_F Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima xylose complex [Thermotoga maritima] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| sp|U6A629|ABFA_PENCN | 1.26e-174 | 12 | 673 | 11 | 630 | Alpha-L-arabinofuranosidase A OS=Penicillium canescens OX=5083 GN=abfA PE=1 SV=1 |
| sp|Q0CTV2|ABFA_ASPTN | 1.28e-106 | 19 | 671 | 26 | 625 | Probable alpha-L-arabinofuranosidase A OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=abfA PE=3 SV=1 |
| sp|A2Q7E0|ABFA_ASPNC | 1.22e-100 | 6 | 671 | 13 | 625 | Probable alpha-L-arabinofuranosidase A OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=abfA PE=3 SV=1 |
| sp|P42254|ABFA_ASPNG | 2.27e-100 | 6 | 671 | 13 | 625 | Alpha-L-arabinofuranosidase A OS=Aspergillus niger OX=5061 GN=abfA PE=1 SV=1 |
| sp|Q8NK90|ABFA_ASPKW | 7.87e-100 | 19 | 671 | 26 | 625 | Alpha-L-arabinofuranosidase A OS=Aspergillus kawachii (strain NBRC 4308) OX=1033177 GN=abfA PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
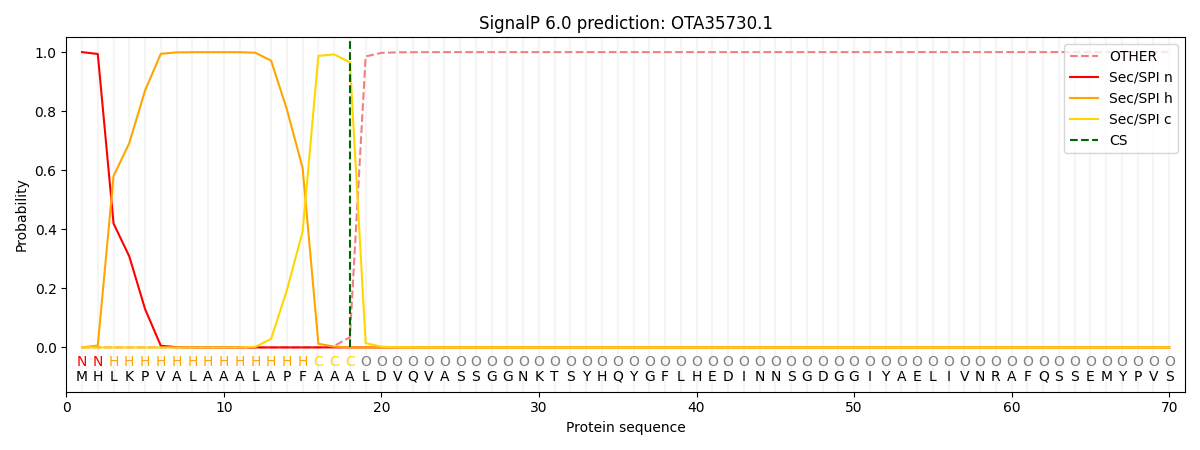
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000250 | 0.999711 | CS pos: 18-19. Pr: 0.9654 |
