You are browsing environment: FUNGIDB
CAZyme Information: OTA30605.1
You are here: Home > Sequence: OTA30605.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Hortaea werneckii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Dothideomycetes; ; Teratosphaeriaceae; Hortaea; Hortaea werneckii | |||||||||||
| CAZyme ID | OTA30605.1 | |||||||||||
| CAZy Family | GH18 | |||||||||||
| CAZyme Description | Alpha-galactosidase [Source:UniProtKB/TrEMBL;Acc:A0A1Z5T3K3] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.22:1 | 3.2.1.88:1 | 3.2.1.22:1 | 3.2.1.88:1 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH27 | 123 | 348 | 4e-51 | 0.8777292576419214 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 269893 | GH27 | 1.56e-66 | 43 | 284 | 1 | 271 | glycosyl hydrolase family 27 (GH27). GH27 enzymes occur in eukaryotes, prokaryotes, and archaea with a wide range of hydrolytic activities, including alpha-glucosidase (glucoamylase and sucrase-isomaltase), alpha-N-acetylgalactosaminidase, and 3-alpha-isomalto-dextranase. All GH27 enzymes cleave a terminal carbohydrate moiety from a substrate that varies considerably in size, depending on the enzyme, and may be either a starch or a glycoprotein. GH27 members are retaining enzymes that cleave their substrates via an acid/base-catalyzed, double-displacement mechanism involving a covalent glycosyl-enzyme intermediate. Two aspartic acid residues have been identified as the catalytic nucleophile and the acid/base, respectively. |
| 166449 | PLN02808 | 4.21e-46 | 40 | 364 | 29 | 376 | alpha-galactosidase |
| 177874 | PLN02229 | 2.94e-43 | 41 | 364 | 61 | 410 | alpha-galactosidase |
| 178295 | PLN02692 | 5.37e-43 | 40 | 364 | 53 | 401 | alpha-galactosidase |
| 271147 | CBM35_galactosidase-like | 2.66e-21 | 386 | 526 | 2 | 125 | Carbohydrate Binding Module family 35 (CBM35); appended mainly to enzymes that bind alpha-D-galactose (CBM35-Gal), including glycoside hydrolase (GH) families GH27 and GH43. This family includes carbohydrate binding module family 35 (CBM35); these are non-catalytic carbohydrate binding domains that are appended mainly to enzymes that bind alpha-D-galactose (CBM35-Gal), including glycoside hydrolase (GH) families GH27 and GH43. Examples of proteins which contain CBM35s belonging to this family includes the CBM35 of an exo-beta-1,3-galactanase from Phanerochaete chrysosporium 9 (Pc1,3Gal43A) which is appended to a GH43 domain, and the CBM35 domain of two bifunctional proteins with beta-L-arabinopyranosidase/alpha-D-galactopyranosidase activities from Fusarium oxysporum 12S, Foap1 and Foap2 (Fo/AP1 and Fo/AP2), that are appended to GH27 domains. CBM35s are unique in that they display conserved specificity through extensive sequence similarity but divergent function through their appended catalytic modules. They are known to bind alpha-D-galactose (Gal), mannan (Man), xylan, glucuronic acid (GlcA), a beta-polymer of mannose, and possibly glucans, forming four subfamilies based on general ligand specificities (galacto, urono, manno, and gluco configurations). Some CBM35s bind their ligands in a calcium-dependent manner. In contrast to most CBMs that are generally rigid proteins, CBM35 undergoes significant conformational change upon ligand binding. GH43 includes beta-xylosidases and beta-xylanases, using aryl-glycosides as substrates, while family GH27 includes alpha-galactosidases, alpha-N-acetylgalactosaminidases, and isomaltodextranases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 5.75e-182 | 40 | 528 | 26 | 561 | |
| 2.49e-178 | 40 | 527 | 26 | 550 | |
| 3.52e-178 | 40 | 527 | 26 | 550 | |
| 4.03e-177 | 40 | 527 | 26 | 550 | |
| 1.15e-176 | 40 | 527 | 26 | 550 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.06e-44 | 40 | 364 | 6 | 352 | Chain A, alpha-galactosidase [Oryza sativa] |
|
| 3.59e-35 | 40 | 346 | 6 | 336 | Nicotiana benthamiana alpha-galactosidase [Nicotiana benthamiana] |
|
| 3.61e-35 | 43 | 369 | 9 | 387 | Crystal structure of alpha-galactosidase I from Mortierella vinacea [Umbelopsis vinacea] |
|
| 1.32e-34 | 40 | 369 | 9 | 408 | Chain A, alpha-galactosidase [Trichoderma reesei],1T0O_A Chain A, alpha-galactosidase [Trichoderma reesei] |
|
| 9.45e-26 | 43 | 361 | 100 | 470 | Crystal structure of a putative alpha-galactosidase (BF1418) from Bacteroides fragilis NCTC 9343 at 1.57 A resolution [Bacteroides fragilis NCTC 9343] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.78e-53 | 41 | 475 | 29 | 591 | Probable alpha-galactosidase D OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=aglD PE=3 SV=2 |
|
| 3.78e-53 | 41 | 475 | 29 | 591 | Probable alpha-galactosidase D OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=aglD PE=3 SV=2 |
|
| 3.21e-50 | 41 | 475 | 29 | 591 | Probable alpha-galactosidase D OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=aglD PE=3 SV=1 |
|
| 2.55e-49 | 43 | 373 | 32 | 403 | Probable alpha-galactosidase D OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=aglD PE=3 SV=2 |
|
| 1.88e-46 | 41 | 475 | 32 | 598 | Probable alpha-galactosidase D OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=aglD PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
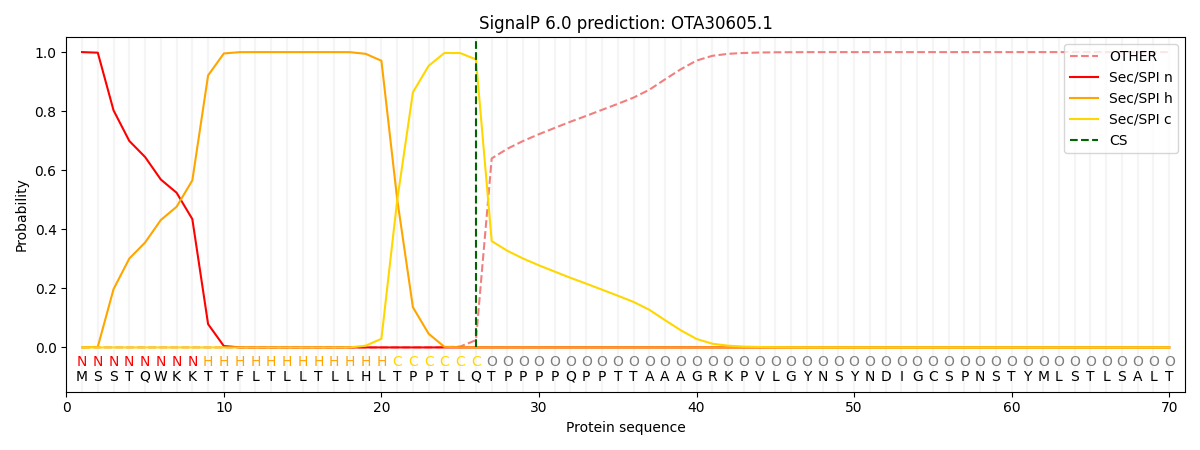
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000281 | 0.999710 | CS pos: 26-27. Pr: 0.9753 |
