You are browsing environment: FUNGIDB
CAZyme Information: ORE11228.1
You are here: Home > Sequence: ORE11228.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Rhizopus microsporus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Mucoromycota; Mucoromycetes; ; Rhizopodaceae; Rhizopus; Rhizopus microsporus | |||||||||||
| CAZyme ID | ORE11228.1 | |||||||||||
| CAZy Family | GT66 | |||||||||||
| CAZyme Description | L-gulonolactone/D-arabinono-1,4-lactone oxidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA7 | 25 | 223 | 4e-25 | 0.4432314410480349 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 273751 | FAD_lactone_ox | 1.62e-162 | 21 | 446 | 2 | 438 | sugar 1,4-lactone oxidases. This model represents a family of at least two different sugar 1,4 lactone oxidases, both involved in synthesizing ascorbic acid or a derivative. These include L-gulonolactone oxidase (EC 1.1.3.8) from rat and D-arabinono-1,4-lactone oxidase (EC 1.1.3.37) from Saccharomyces cerevisiae. Members are proposed to have the cofactor FAD covalently bound at a site specified by Prosite motif PS00862; OX2_COVAL_FAD; 1. |
| 397924 | ALO | 4.64e-114 | 194 | 451 | 2 | 259 | D-arabinono-1,4-lactone oxidase. This domain is specific to D-arabinono-1,4-lactone oxidase EC:1.1.3.-, which is involved in the final step of the D-erythroascorbic acid biosynthesis pathway. |
| 215258 | PLN02465 | 6.13e-63 | 19 | 448 | 82 | 566 | L-galactono-1,4-lactone dehydrogenase |
| 223354 | GlcD | 1.73e-43 | 34 | 452 | 32 | 459 | FAD/FMN-containing dehydrogenase [Energy production and conversion]. |
| 130737 | GLDHase | 8.88e-42 | 19 | 450 | 47 | 538 | galactonolactone dehydrogenase. This model represents L-Galactono-gamma-lactone dehydrogenase (EC 1.3.2.3). This enzyme catalyzes the final step in ascorbic acid biosynthesis in higher plants. This protein is homologous to ascorbic acid biosynthesis enzymes of other species: L-gulono-gamma-lactone oxidase in rat and L-galactono-gamma-lactone oxidase in yeast. All three covalently bind the cofactor FAD. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 2.79e-08 | 16 | 222 | 39 | 259 | |
| 4.91e-08 | 16 | 222 | 39 | 259 | |
| 4.91e-08 | 16 | 222 | 39 | 259 | |
| 1.99e-07 | 42 | 196 | 70 | 227 | |
| 2.55e-07 | 34 | 213 | 48 | 229 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.64e-37 | 19 | 450 | 108 | 602 | Assembly intermediate of the plant mitochondrial complex I [Brassica oleracea] |
|
| 2.00e-35 | 18 | 449 | 5 | 417 | Alditol Oxidase from Streptomyces coelicolor A3(2): Native Enzyme [Streptomyces coelicolor A3(2)],2VFS_A Alditol Oxidase from Streptomyces coelicolor A3(2): Complex with Xylitol [Streptomyces coelicolor A3(2)],2VFT_A Alditol Oxidase from Streptomyces coelicolor A3(2): Complex with Sorbitol [Streptomyces coelicolor A3(2)],2VFU_A Alditol Oxidase from Streptomyces coelicolor A3(2): Complex with Mannitol [Streptomyces coelicolor A3(2)],2VFV_A Alditol Oxidase from Streptomyces coelicolor A3(2): Complex with Sulphite [Streptomyces coelicolor A3(2)] |
|
| 7.67e-08 | 34 | 199 | 206 | 383 | MAMMALIAN ALKYLDIHYDROXYACETONEPHOSPHATE SYNTHASE: Tyr578Phe mutant [Cavia porcellus],4BCA_B MAMMALIAN ALKYLDIHYDROXYACETONEPHOSPHATE SYNTHASE: Tyr578Phe mutant [Cavia porcellus],4BCA_C MAMMALIAN ALKYLDIHYDROXYACETONEPHOSPHATE SYNTHASE: Tyr578Phe mutant [Cavia porcellus],4BCA_D MAMMALIAN ALKYLDIHYDROXYACETONEPHOSPHATE SYNTHASE: Tyr578Phe mutant [Cavia porcellus] |
|
| 7.67e-08 | 34 | 199 | 206 | 383 | MAMMALIAN ALKYLDIHYDROXYACETONEPHOSPHATE SYNTHASE: Arg419His mutant [Cavia porcellus],4BC7_B MAMMALIAN ALKYLDIHYDROXYACETONEPHOSPHATE SYNTHASE: Arg419His mutant [Cavia porcellus],4BC7_C MAMMALIAN ALKYLDIHYDROXYACETONEPHOSPHATE SYNTHASE: Arg419His mutant [Cavia porcellus],4BC7_D MAMMALIAN ALKYLDIHYDROXYACETONEPHOSPHATE SYNTHASE: Arg419His mutant [Cavia porcellus] |
|
| 7.67e-08 | 34 | 199 | 206 | 383 | Mammalian Alkyldihydroxyacetonephosphate Synthase: Wild-Type [Cavia porcellus],4BBY_B Mammalian Alkyldihydroxyacetonephosphate Synthase: Wild-Type [Cavia porcellus],4BBY_C Mammalian Alkyldihydroxyacetonephosphate Synthase: Wild-Type [Cavia porcellus],4BBY_D Mammalian Alkyldihydroxyacetonephosphate Synthase: Wild-Type [Cavia porcellus],4BC9_A MAMMALIAN ALKYLDIHYDROXYACETONEPHOSPHATE SYNTHASE: WILD-TYPE, ADDUCT WITH CYANOETHYL [Cavia porcellus],4BC9_B MAMMALIAN ALKYLDIHYDROXYACETONEPHOSPHATE SYNTHASE: WILD-TYPE, ADDUCT WITH CYANOETHYL [Cavia porcellus],4BC9_C MAMMALIAN ALKYLDIHYDROXYACETONEPHOSPHATE SYNTHASE: WILD-TYPE, ADDUCT WITH CYANOETHYL [Cavia porcellus],4BC9_D MAMMALIAN ALKYLDIHYDROXYACETONEPHOSPHATE SYNTHASE: WILD-TYPE, ADDUCT WITH CYANOETHYL [Cavia porcellus],5ADZ_A Ether Lipid-Generating Enzyme AGPS in complex with inhibitor 1a [Cavia porcellus],5ADZ_B Ether Lipid-Generating Enzyme AGPS in complex with inhibitor 1a [Cavia porcellus],5ADZ_C Ether Lipid-Generating Enzyme AGPS in complex with inhibitor 1a [Cavia porcellus],5ADZ_D Ether Lipid-Generating Enzyme AGPS in complex with inhibitor 1a [Cavia porcellus],5AE1_A Ether Lipid-Generating Enzyme AGPS in complex with inhibitor ZINC69435460 [Cavia porcellus],5AE1_B Ether Lipid-Generating Enzyme AGPS in complex with inhibitor ZINC69435460 [Cavia porcellus],5AE1_C Ether Lipid-Generating Enzyme AGPS in complex with inhibitor ZINC69435460 [Cavia porcellus],5AE1_D Ether Lipid-Generating Enzyme AGPS in complex with inhibitor ZINC69435460 [Cavia porcellus],5AE2_A Ether Lipid-Generating Enzyme AGPS in complex with inhibitor 1e [Cavia porcellus],5AE2_B Ether Lipid-Generating Enzyme AGPS in complex with inhibitor 1e [Cavia porcellus],5AE2_C Ether Lipid-Generating Enzyme AGPS in complex with inhibitor 1e [Cavia porcellus],5AE2_D Ether Lipid-Generating Enzyme AGPS in complex with inhibitor 1e [Cavia porcellus],5AE3_A Ether Lipid-Generating Enzyme AGPS in complex with antimycin A [Cavia porcellus],5AE3_B Ether Lipid-Generating Enzyme AGPS in complex with antimycin A [Cavia porcellus],5AE3_C Ether Lipid-Generating Enzyme AGPS in complex with antimycin A [Cavia porcellus],5AE3_D Ether Lipid-Generating Enzyme AGPS in complex with antimycin A [Cavia porcellus],6GOU_A Development of Alkyl Glycerone Phosphate Synthase Inhibitors: Complex with Inhibitor 2I [Cavia porcellus],6GOU_B Development of Alkyl Glycerone Phosphate Synthase Inhibitors: Complex with Inhibitor 2I [Cavia porcellus],6GOU_C Development of Alkyl Glycerone Phosphate Synthase Inhibitors: Complex with Inhibitor 2I [Cavia porcellus],6GOU_D Development of Alkyl Glycerone Phosphate Synthase Inhibitors: Complex with Inhibitor 2I [Cavia porcellus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.25e-134 | 21 | 450 | 8 | 437 | L-gulonolactone oxidase OS=Scyliorhinus torazame OX=75743 GN=GULO PE=2 SV=1 |
|
| 5.03e-126 | 23 | 450 | 10 | 437 | L-gulonolactone oxidase OS=Rattus norvegicus OX=10116 GN=Gulo PE=1 SV=3 |
|
| 2.86e-125 | 23 | 450 | 10 | 437 | L-gulonolactone oxidase OS=Mus musculus OX=10090 GN=Gulo PE=1 SV=3 |
|
| 1.15e-124 | 23 | 450 | 10 | 437 | L-gulonolactone oxidase OS=Bos taurus OX=9913 GN=GULO PE=2 SV=3 |
|
| 2.30e-124 | 23 | 450 | 10 | 437 | L-gulonolactone oxidase OS=Sus scrofa OX=9823 GN=GULO PE=2 SV=3 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
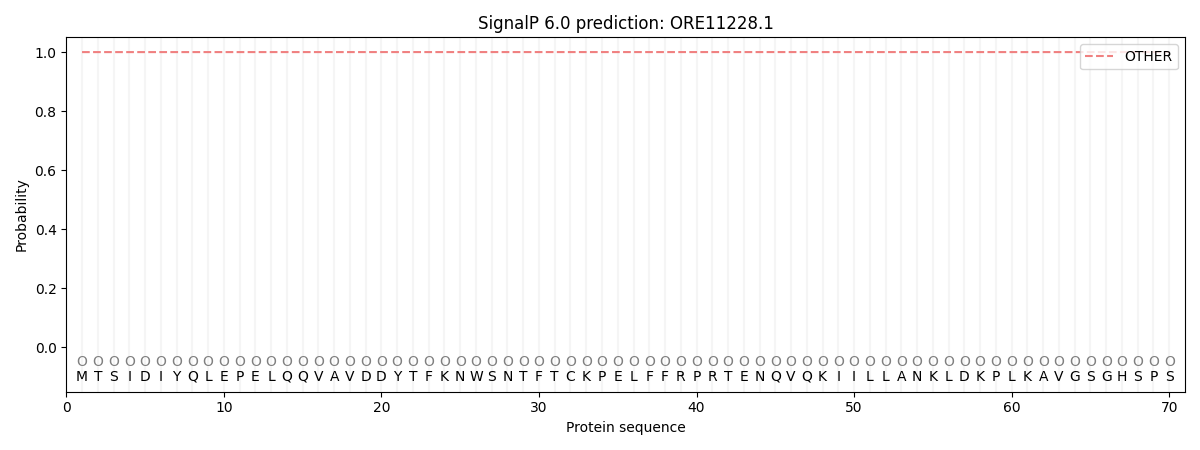
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000065 | 0.000000 |
