You are browsing environment: FUNGIDB
CAZyme Information: ORE06335.1
You are here: Home > Sequence: ORE06335.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
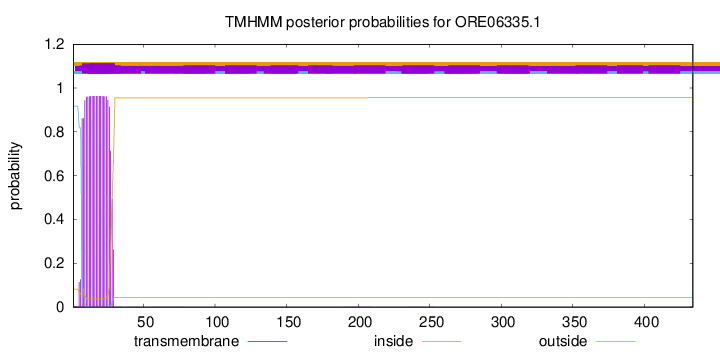
TMHMM annotations
Basic Information help
| Species | Rhizopus microsporus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Mucoromycota; Mucoromycetes; ; Rhizopodaceae; Rhizopus; Rhizopus microsporus | |||||||||||
| CAZyme ID | ORE06335.1 | |||||||||||
| CAZy Family | GH47|GH47 | |||||||||||
| CAZyme Description | glycosyl transferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 2.4.1.131:20 | 2.4.1.-:1 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT15 | 82 | 359 | 4.2e-111 | 0.9926739926739927 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 227353 | KTR1 | 1.28e-140 | 81 | 382 | 79 | 375 | Mannosyltransferase [Carbohydrate transport and metabolism]. |
| 396385 | Glyco_transf_15 | 7.51e-120 | 73 | 358 | 33 | 311 | Glycolipid 2-alpha-mannosyltransferase. This is a family of alpha-1,2 mannosyl-transferases involved in N-linked and O-linked glycosylation of proteins. Some of the enzymes in this family have been shown to be involved in O- and N-linked glycan modifications in the Golgi. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.91e-170 | 58 | 431 | 58 | 412 | |
| 4.09e-126 | 72 | 378 | 54 | 358 | |
| 6.64e-118 | 54 | 416 | 46 | 374 | |
| 8.80e-118 | 57 | 387 | 41 | 361 | |
| 1.33e-117 | 82 | 384 | 57 | 358 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.76e-112 | 74 | 417 | 18 | 335 | Crystal structure of yeast alpha1,2-mannosyltransferase Kre2p/Mnt1p [Saccharomyces cerevisiae],1S4N_B Crystal structure of yeast alpha1,2-mannosyltransferase Kre2p/Mnt1p [Saccharomyces cerevisiae],1S4O_A Crystal structure of yeast alpha1,2-mannosyltransferase Kre2p/Mnt1p: binary complex with GDP/Mn [Saccharomyces cerevisiae],1S4O_B Crystal structure of yeast alpha1,2-mannosyltransferase Kre2p/Mnt1p: binary complex with GDP/Mn [Saccharomyces cerevisiae],1S4P_A Crystal structure of yeast alpha1,2-mannosyltransferase Kre2p/Mnt1p: ternary complex with GDP/Mn and methyl-alpha-mannoside acceptor [Saccharomyces cerevisiae],1S4P_B Crystal structure of yeast alpha1,2-mannosyltransferase Kre2p/Mnt1p: ternary complex with GDP/Mn and methyl-alpha-mannoside acceptor [Saccharomyces cerevisiae] |
|
| 1.60e-79 | 75 | 378 | 59 | 360 | Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, apo form [Aspergillus fumigatus A1163],7BOO_B Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, apo form [Aspergillus fumigatus A1163],7BOO_C Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, apo form [Aspergillus fumigatus A1163],7BOO_D Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, apo form [Aspergillus fumigatus A1163],7BOO_E Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, apo form [Aspergillus fumigatus A1163],7BOO_F Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, apo form [Aspergillus fumigatus A1163],7BOP_A Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, Mn/GDP-form [Aspergillus fumigatus A1163],7BOP_B Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, Mn/GDP-form [Aspergillus fumigatus A1163],7BOP_C Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, Mn/GDP-form [Aspergillus fumigatus A1163],7BOP_D Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, Mn/GDP-form [Aspergillus fumigatus A1163],7BOP_E Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, Mn/GDP-form [Aspergillus fumigatus A1163],7BOP_F Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, Mn/GDP-form [Aspergillus fumigatus A1163] |
|
| 5.88e-68 | 82 | 365 | 102 | 388 | X-ray structure of the mannosyltransferase Ktr4p from S. cerevisiae in complex with GDP [Saccharomyces cerevisiae S288C],5A07_B X-ray structure of the mannosyltransferase Ktr4p from S. cerevisiae in complex with GDP [Saccharomyces cerevisiae S288C],5A08_A X-ray structure of the mannosyltransferase Ktr4p from S. cerevisiae [Saccharomyces cerevisiae S288C],5A08_B X-ray structure of the mannosyltransferase Ktr4p from S. cerevisiae [Saccharomyces cerevisiae S288C] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.53e-117 | 81 | 378 | 138 | 430 | Glycolipid 2-alpha-mannosyltransferase 2 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=MNT2 PE=3 SV=4 |
|
| 2.02e-116 | 79 | 387 | 67 | 370 | Alpha-1,2 mannosyltransferase KTR1 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=KTR1 PE=1 SV=1 |
|
| 7.10e-113 | 73 | 378 | 99 | 399 | Glycolipid 2-alpha-mannosyltransferase 1 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=MNT1 PE=3 SV=1 |
|
| 1.03e-109 | 74 | 417 | 112 | 429 | Glycolipid 2-alpha-mannosyltransferase OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=KRE2 PE=1 SV=1 |
|
| 2.10e-104 | 81 | 378 | 68 | 358 | O-glycoside alpha-1,2-mannosyltransferase omh1 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=omh1 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
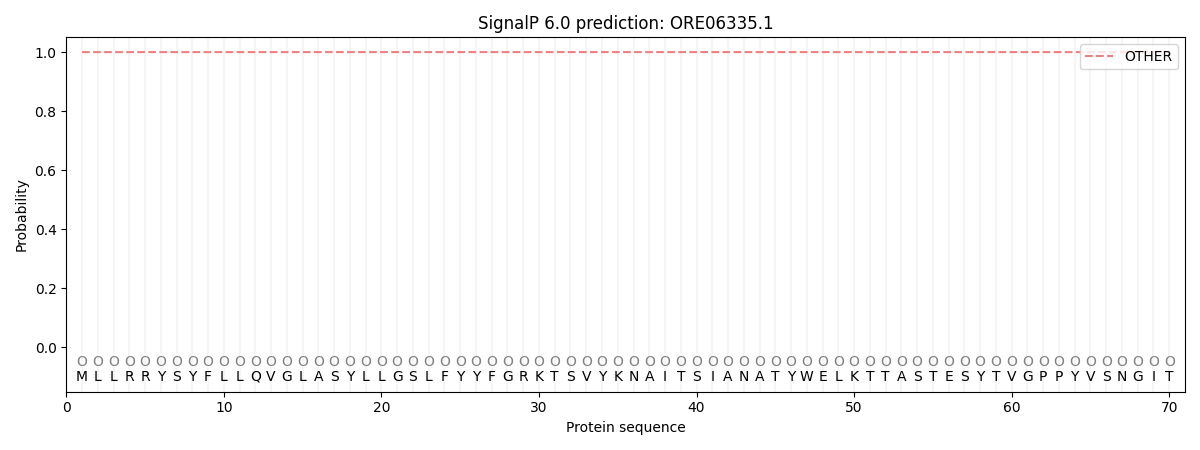
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000027 | 0.000005 |