You are browsing environment: FUNGIDB
CAZyme Information: OJD23674.1
You are here: Home > Sequence: OJD23674.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Blastomyces percursus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Ajellomycetaceae; Blastomyces; Blastomyces percursus | |||||||||||
| CAZyme ID | OJD23674.1 | |||||||||||
| CAZy Family | GH43 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH17 | 25 | 302 | 4.8e-41 | 0.9935691318327974 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 227625 | Scw11 | 3.54e-71 | 8 | 294 | 30 | 305 | Exo-beta-1,3-glucanase, GH17 family [Carbohydrate transport and metabolism]. |
| 366033 | Glyco_hydro_17 | 4.82e-51 | 25 | 302 | 2 | 309 | Glycosyl hydrolases family 17. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 7.59e-193 | 1 | 302 | 1 | 302 | |
| 1.26e-191 | 1 | 302 | 1 | 302 | |
| 3.60e-191 | 1 | 302 | 1 | 302 | |
| 3.43e-160 | 1 | 302 | 1 | 301 | |
| 4.64e-158 | 1 | 302 | 1 | 301 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.97e-38 | 24 | 297 | 38 | 289 | Crystal structure of glycoside hydrolase family 17 beta-1,3-glucanosyltransferase from Rhizomucor miehei [Rhizomucor miehei CAU432] |
|
| 1.07e-37 | 24 | 297 | 38 | 289 | Active-site mutant of Rhizomucor miehei beta-1,3-glucanosyltransferase in complex with laminaribiose [Rhizomucor miehei CAU432],4WTS_A Active-site mutant of Rhizomucor miehei beta-1,3-glucanosyltransferase in complex with laminaritriose [Rhizomucor miehei CAU432] |
|
| 3.01e-07 | 55 | 298 | 101 | 381 | Crystal structure of an endo-laminarinase from Formosa Hel1_33_131 [Formosa sp. Hel1_33_131],6FCG_B Crystal structure of an endo-laminarinase from Formosa Hel1_33_131 [Formosa sp. Hel1_33_131],6FCG_C Crystal structure of an endo-laminarinase from Formosa Hel1_33_131 [Formosa sp. Hel1_33_131],6FCG_D Crystal structure of an endo-laminarinase from Formosa Hel1_33_131 [Formosa sp. Hel1_33_131],6FCG_E Crystal structure of an endo-laminarinase from Formosa Hel1_33_131 [Formosa sp. Hel1_33_131],6FCG_F Crystal structure of an endo-laminarinase from Formosa Hel1_33_131 [Formosa sp. Hel1_33_131] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.76e-146 | 1 | 302 | 1 | 301 | Glucan 1,3-beta-glucosidase ARB_02797 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_02797 PE=1 SV=1 |
|
| 8.22e-76 | 18 | 302 | 16 | 305 | Glucan 1,3-beta-glucosidase BGL2 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=BGL2 PE=1 SV=2 |
|
| 1.87e-74 | 18 | 302 | 16 | 305 | Glucan 1,3-beta-glucosidase OS=Candida albicans OX=5476 GN=BGL2 PE=3 SV=1 |
|
| 4.92e-73 | 18 | 302 | 21 | 310 | Glucan 1,3-beta-glucosidase OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=BGL2 PE=1 SV=1 |
|
| 2.03e-62 | 19 | 302 | 37 | 318 | Glucan 1,3-beta-glucosidase OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=bgl2 PE=2 SV=4 |
SignalP and Lipop Annotations help
This protein is predicted as SP
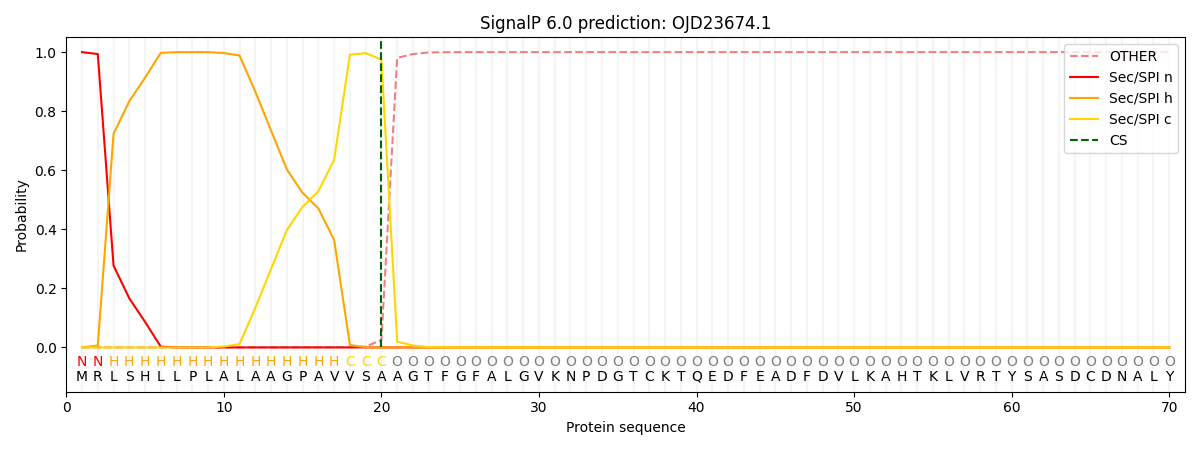
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000237 | 0.999725 | CS pos: 20-21. Pr: 0.9748 |

