You are browsing environment: FUNGIDB
CAZyme Information: OJD14025.1
You are here: Home > Sequence: OJD14025.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Emergomyces pasteurianus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Ajellomycetaceae; Emergomyces; Emergomyces pasteurianus | |||||||||||
| CAZyme ID | OJD14025.1 | |||||||||||
| CAZy Family | GH18 | |||||||||||
| CAZyme Description | FAD-binding PCMH-type domain-containing protein [Source:UniProtKB/TrEMBL;Acc:A0A1J9PBY3] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA7 | 131 | 322 | 2.7e-41 | 0.3930131004366812 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 223354 | GlcD | 1.73e-18 | 116 | 569 | 18 | 449 | FAD/FMN-containing dehydrogenase [Energy production and conversion]. |
| 396238 | FAD_binding_4 | 4.48e-16 | 131 | 278 | 1 | 139 | FAD binding domain. This family consists of various enzymes that use FAD as a co-factor, most of the enzymes are similar to oxygen oxidoreductase. One of the enzymes Vanillyl-alcohol oxidase (VAO) has a solved structure, the alignment includes the FAD binding site, called the PP-loop, between residues 99-110. The FAD molecule is covalently bound in the known structure, however the residue that links to the FAD is not in the alignment. VAO catalyzes the oxidation of a wide variety of substrates, ranging form aromatic amines to 4-alkylphenols. Other members of this family include D-lactate dehydrogenase, this enzyme catalyzes the conversion of D-lactate to pyruvate using FAD as a co-factor; mitomycin radical oxidase, this enzyme oxidizes the reduced form of mitomycins and is involved in mitomycin resistance. This family includes MurB an UDP-N-acetylenolpyruvoylglucosamine reductase enzyme EC:1.1.1.158. This enzyme is involved in the biosynthesis of peptidoglycan. |
| 369658 | BBE | 5.26e-08 | 532 | 574 | 1 | 42 | Berberine and berberine like. This domain is found in the berberine bridge and berberine bridge- like enzymes which are involved in the biosynthesis of numerous isoquinoline alkaloids. They catalyze the transformation of the N-methyl group of (S)-reticuline into the C-8 berberine bridge carbon of (S)-scoulerine. |
| 178402 | PLN02805 | 8.71e-07 | 129 | 333 | 132 | 333 | D-lactate dehydrogenase [cytochrome] |
| 215242 | PLN02441 | 3.56e-05 | 255 | 316 | 187 | 241 | cytokinin dehydrogenase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.57e-39 | 13 | 586 | 493 | 1059 | |
| 1.57e-39 | 13 | 586 | 493 | 1059 | |
| 6.10e-15 | 131 | 578 | 63 | 491 | |
| 1.10e-14 | 131 | 578 | 70 | 497 | |
| 1.44e-14 | 138 | 577 | 72 | 490 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.08e-63 | 30 | 579 | 39 | 561 | Crystal structure of VAO-type flavoprotein MtVAO615 at pH 7.5 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464],6F73_A Crystal structure of VAO-type flavoprotein MtVAO615 at pH 5.0 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464],6F73_B Crystal structure of VAO-type flavoprotein MtVAO615 at pH 5.0 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464] |
|
| 1.00e-53 | 34 | 579 | 45 | 585 | Crystal structure of VAO-type flavoprotein MtVAO713 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464],6F74_B Crystal structure of VAO-type flavoprotein MtVAO713 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464],6F74_C Crystal structure of VAO-type flavoprotein MtVAO713 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464],6F74_D Crystal structure of VAO-type flavoprotein MtVAO713 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464] |
|
| 1.24e-16 | 131 | 340 | 53 | 249 | Phl p 4 I153V N158H variant, a glucose oxidase [Phleum pratense],4PWC_A Phl p 4 I153V N158H variant, a glucose oxidase, 3.5 M NaBr soak [Phleum pratense] |
|
| 1.64e-16 | 131 | 340 | 53 | 249 | Phl p 4 N158H variant, a glucose dehydrogenase [Phleum pratense] |
|
| 2.67e-16 | 131 | 576 | 48 | 468 | Physcomitrella patens BBE-like 1 variant D396N [Physcomitrium patens],6EO5_B Physcomitrella patens BBE-like 1 variant D396N [Physcomitrium patens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.19e-94 | 21 | 579 | 13 | 589 | FAD-linked oxidoreductase ffsJ OS=Aspergillus flavipes OX=41900 GN=ffsJ PE=3 SV=1 |
|
| 2.52e-83 | 30 | 577 | 25 | 562 | FAD-linked oxidoreductase OXR2 OS=Magnaporthe oryzae (strain 70-15 / ATCC MYA-4617 / FGSC 8958) OX=242507 GN=OXR2 PE=2 SV=1 |
|
| 1.42e-75 | 19 | 463 | 12 | 421 | FAD-linked oxidoreductase phmC OS=Phaeosphaeria nodorum (strain SN15 / ATCC MYA-4574 / FGSC 10173) OX=321614 GN=phmC PE=3 SV=2 |
|
| 8.85e-69 | 30 | 579 | 65 | 596 | FAD-linked oxidoreductase cheF OS=Chaetomium globosum (strain ATCC 6205 / CBS 148.51 / DSM 1962 / NBRC 6347 / NRRL 1970) OX=306901 GN=cheF PE=2 SV=1 |
|
| 3.24e-65 | 30 | 579 | 38 | 550 | FAD-linked oxidoreductase ZEB1 OS=Gibberella zeae (strain ATCC MYA-4620 / CBS 123657 / FGSC 9075 / NRRL 31084 / PH-1) OX=229533 GN=ZEB1 PE=2 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
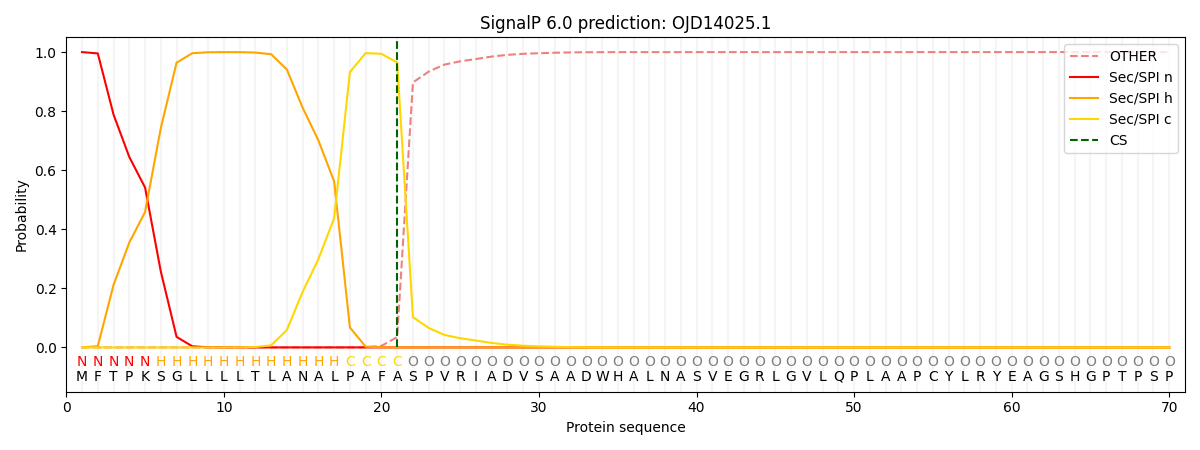
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000535 | 0.999434 | CS pos: 21-22. Pr: 0.9649 |
