You are browsing environment: FUNGIDB
CAZyme Information: OJD13280.1
You are here: Home > Sequence: OJD13280.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
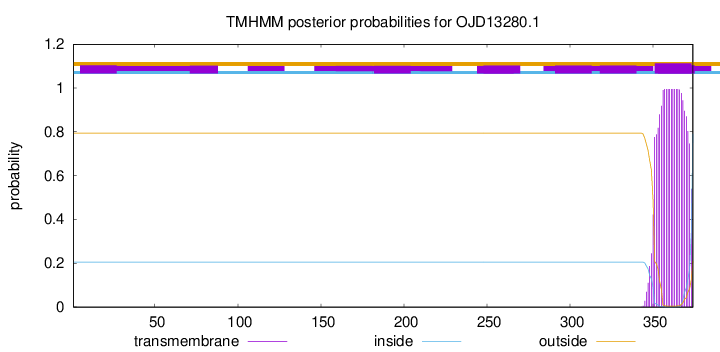
TMHMM annotations
Basic Information help
| Species | Emergomyces pasteurianus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Ajellomycetaceae; Emergomyces; Emergomyces pasteurianus | |||||||||||
| CAZyme ID | OJD13280.1 | |||||||||||
| CAZy Family | GH16 | |||||||||||
| CAZyme Description | 1,3-beta-glucanosyltransferase [Source:UniProtKB/TrEMBL;Acc:A0A1J9P9W2] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH72 | 1 | 231 | 1.4e-91 | 0.7403846153846154 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 397351 | Glyco_hydro_72 | 2.57e-98 | 2 | 267 | 64 | 312 | Glucanosyltransferase. This is a family of glycosylphosphatidylinositol-anchored beta(1-3)glucanosyltransferases. The active site residues in the Aspergillus fumigatus example are the two glutamate residues at 160 and 261. |
| 223033 | PHA03291 | 2.23e-04 | 240 | 340 | 160 | 276 | envelope glycoprotein I; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.74e-226 | 1 | 374 | 1 | 378 | |
| 1.93e-223 | 1 | 374 | 1 | 378 | |
| 3.36e-198 | 1 | 303 | 152 | 454 | |
| 3.33e-153 | 1 | 374 | 72 | 447 | |
| 8.42e-152 | 1 | 318 | 72 | 390 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.01e-65 | 1 | 299 | 96 | 375 | Saccharomyces cerevisiae Gas2p in complex with laminaripentaose [Saccharomyces cerevisiae],2W63_A Saccharomyces Cerevisiae Gas2p In Complex With Laminaritriose And Laminaritetraose [Saccharomyces cerevisiae],5O9O_A Crystal structure of ScGas2 in complex with compound 7. [Saccharomyces cerevisiae S288C],5O9P_A Crystal structure of Gas2 in complex with compound 10 [Saccharomyces cerevisiae S288C],5O9Q_A Crystal structure of ScGas2 in complex with compound 6 [Saccharomyces cerevisiae S288C],5O9R_A Crystal structure of ScGas2 in complex with compound 9 [Saccharomyces cerevisiae S288C],5O9Y_A Crystal structure of ScGas2 in complex with compound 11 [Saccharomyces cerevisiae S288C],5OA2_A Crystal structure of ScGas2 in complex with compound 8 [Saccharomyces cerevisiae S288C],5OA2_B Crystal structure of ScGas2 in complex with compound 8 [Saccharomyces cerevisiae S288C],5OA2_C Crystal structure of ScGas2 in complex with compound 8 [Saccharomyces cerevisiae S288C],5OA6_A Crystal structure of ScGas2 in complex with compound 12 [Saccharomyces cerevisiae S288C] |
|
| 2.78e-65 | 1 | 299 | 96 | 375 | Saccharomyces cerevisiae Gas2p apostructure (E176Q mutant) [Saccharomyces cerevisiae] |
|
| 2.78e-65 | 1 | 299 | 96 | 375 | SACCHAROMYCES CEREVISIAE GAS2P (E176Q MUTANT) IN COMPLEX WITH LAMINARITETRAOSE AND LAMINARIPENTAOSE [Saccharomyces cerevisiae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.21e-73 | 1 | 289 | 75 | 344 | 1,3-beta-glucanosyltransferase gel3 OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=gel3 PE=3 SV=1 |
|
| 9.87e-73 | 1 | 289 | 75 | 344 | 1,3-beta-glucanosyltransferase gel3 OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=gel3 PE=1 SV=1 |
|
| 4.55e-69 | 1 | 305 | 76 | 364 | 1,3-beta-glucanosyltransferase gas2 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=gas2 PE=2 SV=1 |
|
| 3.02e-66 | 1 | 299 | 89 | 366 | pH-responsive protein 1 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=PHR1 PE=2 SV=4 |
|
| 3.58e-66 | 1 | 278 | 113 | 381 | 1,3-beta-glucanosyltransferase PGA5 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=PGA5 PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
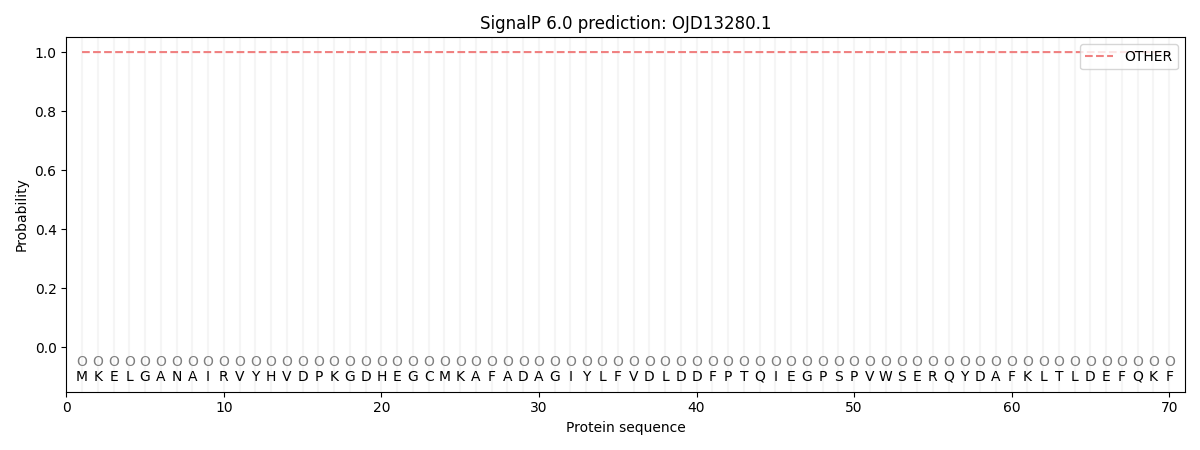
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000066 | 0.000000 |